THE FUTURE OF
COMPLEX BIOLOGY
The CellXpress.ai Automated Cell Culture System
See the difference. Pick the best. QPix FLEX Microbial Colony Picking System
Dive Deeper: ImageXpress HCS.ai High-Content Screening System with AI-powered Insights
Industry-leading SpectraMax® Microplate Readers and SoftMax® Pro Software
A life science company
We journey alongside researchers, combining next-gen technology and deep expertise to spark bold breakthroughs that push research boundaries and transform lives worldwide.
Advancing scientific discovery
Unlocking the full potential of complex biology, human-relevant models, AI-driven discovery, and next-generation therapeutics is key to navigating what's next in life sciences. We partner with you to pave the way.


We believe in the power of collaboration.
Today’s scientific breakthroughs demand more than just advanced tools. They require partnerships built on understanding, commitment, and trust. Collaboration in life sciences plays a critical role in fueling advancements and unlocking new frontiers in discovery.
At Molecular Devices, we stand shoulder to shoulder with customers as indispensable partners. Whether enabling the adoption of complex biology, supporting human-relevant model research, integrating AI and machine learning, or validating the next generation of therapeutic modalities, we empower transformational breakthroughs.

Unlocking the full potential of complex biology to revolutionize research.
Our industry expertise and advanced technologies are unlocking the potential for complex biology to revolutionize scientific research. We partner with customers to create tailored, automated workflows that streamline complexity, increase throughput, and accelerate breakthroughs with confidence.

Making human-relevant model-based research more accessible.
Complex cell models mirroring human biology are key to reducing the persistently high attrition rate of drug candidates in clinical trials. From automated cell culture systems and screening solutions to an abundance of ready-for-use organoids from our lines or yours, we enable researchers to reliably assess the safety and efficacy of therapies based on human-relevant models from the outset.
Unleashing the power of AI and machine learning to advance discovery.

Harnessing AI and machine learning puts scientists in the strongest ever position to industrialize and interrogate complex biology. From cell culturing to smart imaging to advanced data analysis, our automated, AI-enabled, end-to-end solutions free researchers from making regular, mundane checks so they can invest their time on transformative science.
Assuring validation to accelerate time to market for new therapeutics

If new therapeutic modalities are to realize their promise, traceable audit-ready software and hardware must evolve alongside them. Our solutions offer the data integrity needed to accelerate next-generation therapies.
Your indispensable partner in complex biology
Our advanced instruments and automated workflows—built alongside you—unlock new possibilities, streamlining complexity so you can focus on discovery.
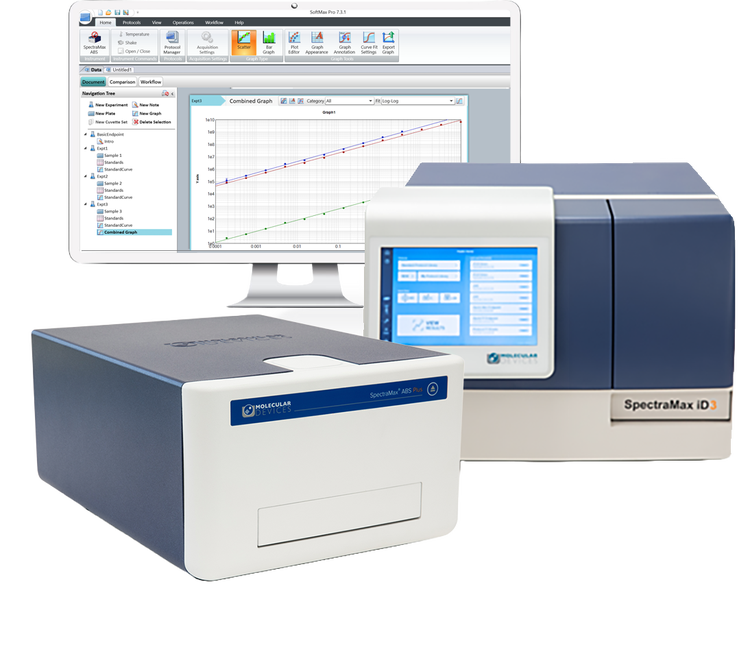

Easy-to-use, intuitive, configurable microplate readers
For over 40 years, we’ve partnered with scientists to expand the boundaries of their research. Our SpectraMax microplate readers and SoftMax Pro software are the industry's most cited and have empowered life science researchers to advance protein and cell biology―breaking barriers to novel, landmark discoveries.
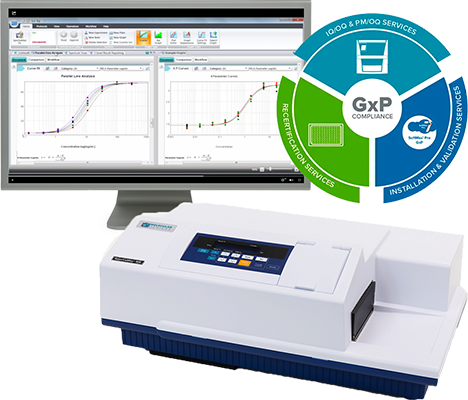

Streamline your GxP compliance journey for GMP/GLP labs
A leader in comprehensive GxP compliance solutions, we combine our SpectraMax® microplate readers, SoftMax® Pro GxP software, SpectraTest® Validation Plates and expert IQ/OQ/PM services to help laboratories operating under GMP and GLP regulations follow FDA or regional regulatory guidelines with confidence.
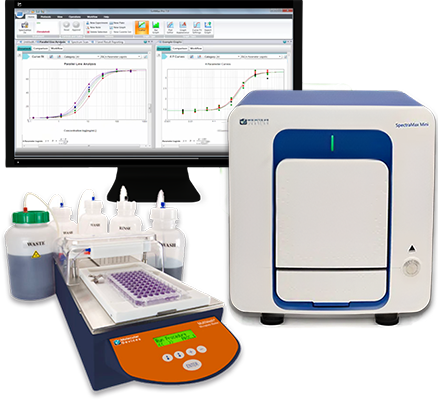
Save on a bundle: Microplate reader and washer
Improve your lab’s efficiency, get more data, and decrease time to discovery with an automated microplate reader and washer solution:
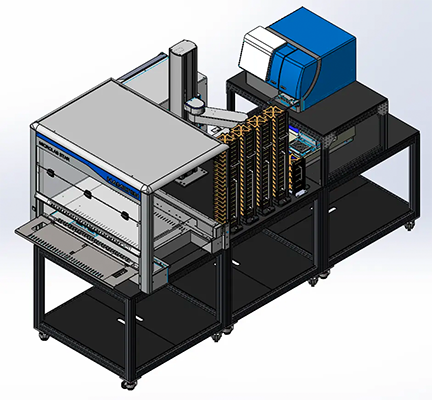
Lab automation for high-throughput plate-based assays
Explore our fully-integrated, automated workflow solutions for cell and biochemical assays. Our scalable ELISA Workcells provide walkaway time, increasing throughput, effectiveness and efficiency of the assay procedure, and reproducibility.
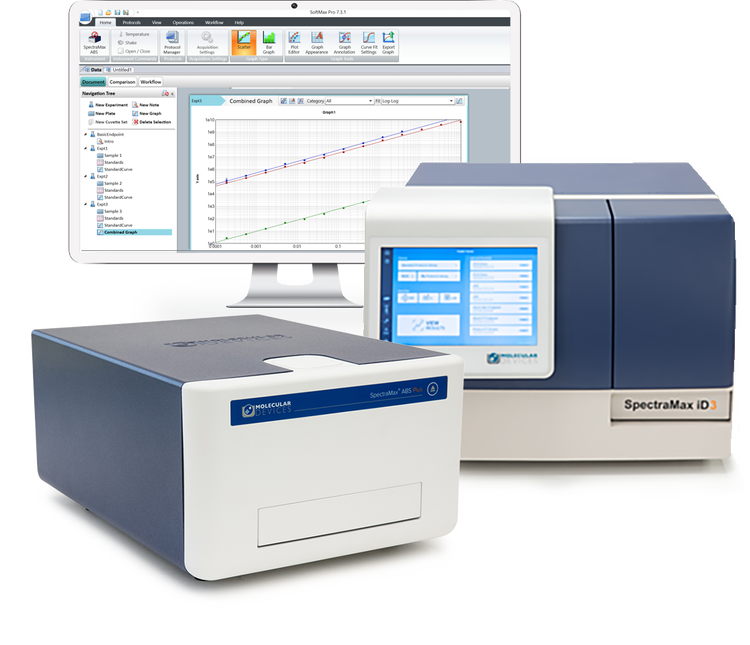

Easy-to-use, intuitive, configurable microplate readers
For over 40 years, we’ve partnered with scientists to expand the boundaries of their research. Our SpectraMax microplate readers and SoftMax Pro software are the industry's most cited and have empowered life science researchers to advance protein and cell biology―breaking barriers to novel, landmark discoveries.

Streamline your GxP compliance journey for GMP/GLP labs
A leader in comprehensive GxP compliance solutions, we combine our SpectraMax® microplate readers, SoftMax® Pro GxP software, SpectraTest® Validation Plates and expert IQ/OQ/PM services to help laboratories operating under GMP and GLP regulations follow FDA or regional regulatory guidelines with confidence.

Lab automation for high-throughput plate-based assays
Explore our fully-integrated, automated workflow solutions for cell and biochemical assays. Our scalable ELISA Workcells provide walkaway time, increasing throughput, effectiveness and efficiency of the assay procedure, and reproducibility.


ImageXpress® HCS.ai High-Content Screening System for superior data analysis and AI-driven insights
Distinguished by its advanced, modular design and ability to quickly capture crystal-clear images of complex cell models, acquire detailed data with intuitive software, and offer deep insights leveraging AI-driven analysis.


Introducing the revolutionary CellXpress.ai® Automated Cell Culture System
An Al-driven cell culture innovation hub, the CellXpress.ai system automates processes, improves workflows, and makes assays more reliable and reproducible with machine learning-assisted monitoring, feeding, imaging and scheduling.
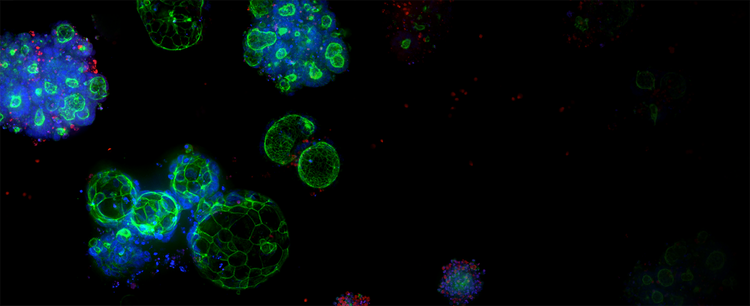
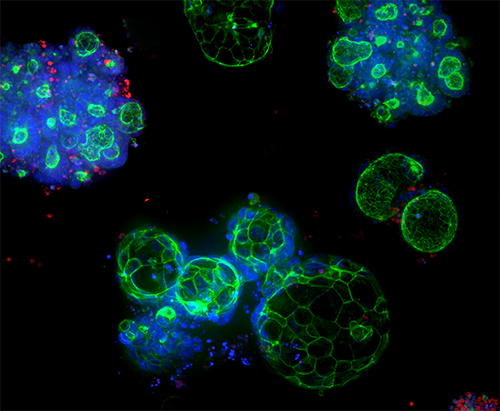
3D Ready Organoids and Organoid Expansion Services
Quality-controlled organoids are manufactured at scale for high-throughput screening, leveraging proprietary bioreactor and bioprocess technology to produce reliable and predictive patient-derived organoids (PDOs).

The QPix® FLEX™ Microbial Colony Picking System for precision, flexibility, and space-limited environments
Plating, streaking, picking, and liquid handling, all in one compact device? The QPix FLEX Microbial Colony Picker makes it look easy. With advanced color imaging, it helps you spot the right colonies faster. And yes, it fits on your bench or in a hypoxic chamber – because why wouldn’t it? The QPix FLEX system replaces the clutter with one smart, streamlined system built to move your research forward.
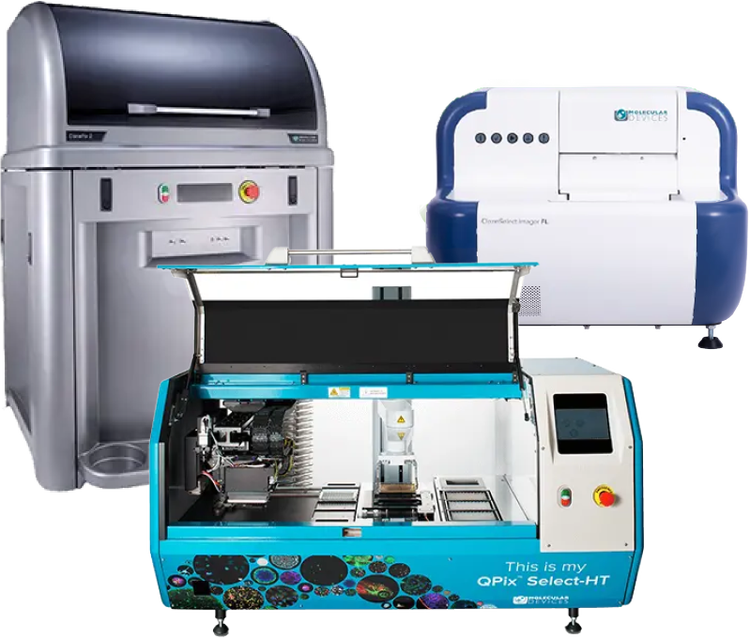

Clone screening systems for colony picking and single-cell isolation
The QPix® 400 Microbial Colony Picker, ClonePix® 2 Mammalian Colony Picker, and CloneSelect® FL Imager increase throughput and consistency across cell line development, monoclonal antibody discovery, and synthetic biology workflows.

Automated solutions for high-throughput clone screening
Fully-integrated, lab automation solution for molecular cloning, antibody discovery, and monoclonality. Our automated clone screening workflows integrate laboratory devices to increase your throughput and efficiency while reducing human interaction.

Single-cell dispensing and screening for monoclonality verification.
Award-winning DispenCell™ Single-Cell Dispenser and the CloneSelect® Imager FL – our bundled solution offers unparalleled precision and efficiency in cell line development and monoclonality verification at day zero.
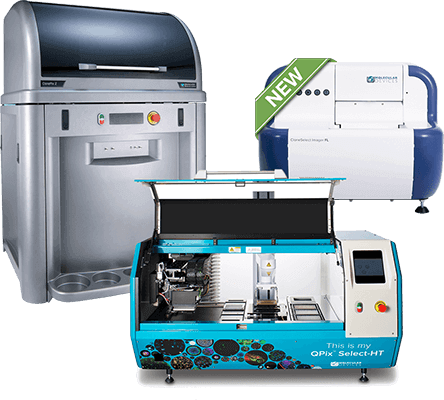

Clone screening systems for colony picking and single-cell isolation
The QPix® 400 Microbial Colony Picker, ClonePix® 2 Mammalian Colony Picker, and CloneSelect® FL Imager increase throughput and consistency across cell line development, monoclonal antibody discovery, and synthetic biology workflows.

Automated solutions for high-throughput clone screening
Fully-integrated, lab automation solution for molecular cloning, antibody discovery, and monoclonality. Our automated clone screening workflows integrate laboratory devices to increase your throughput and efficiency while reducing human interaction.
We’re scientists supporting scientists
Our experts help researchers move beyond traditional testing with complex, human-relevant models—unlocking faster, smarter breakthroughs that truly translate.

3D Biology: The paradigm shift in next-gen drug discovery
In the midst of an evolving drug discovery paradigm, researchers worldwide are transitioning their compound screens away from 2D cell cultures and animal models to more complex, human-relevant 3D systems like organoids. Experience our interactive infographic as it takes you deeper into why the industry is embracing this next generation of drug discovery and the innovations supporting scientists in their 3D biology journey.
Drug Discovery & Development
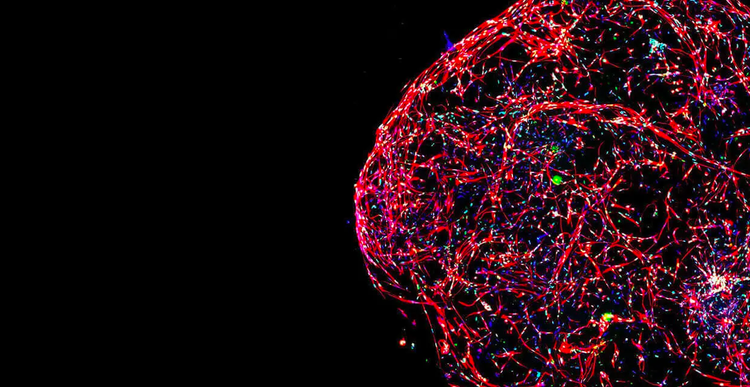
The drug discovery landscape is shifting, with more scientists centering cell line development, disease models, and high-throughput screening methods around physiologically-relevant 3D cell models.
The reason for this is clear: Using cellular model systems in research that closely mimic patient disease states or human organs can bring life-saving therapeutics to market – faster.
3D Cell Models
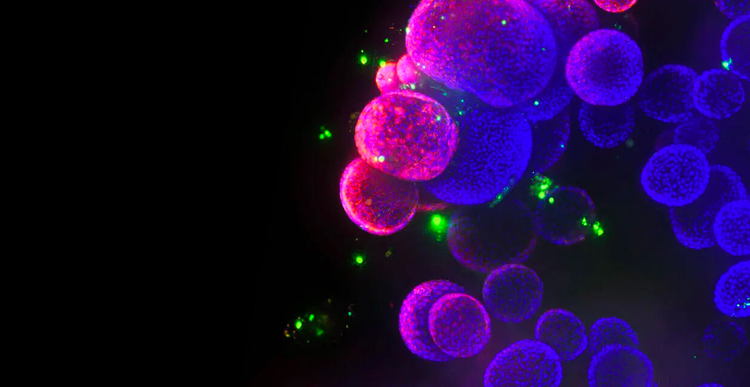
With our automated 3D cell culture and bioimaging screening solutions, we are helping reshape the future of drug discovery. Supported by our technology and organoid development protocols, researchers can now advance and scale screening methods for physiologically-relevant 3D models that more closely mimic patient disease states and human organs, leading to faster drug development and approval.
Cell Line Development
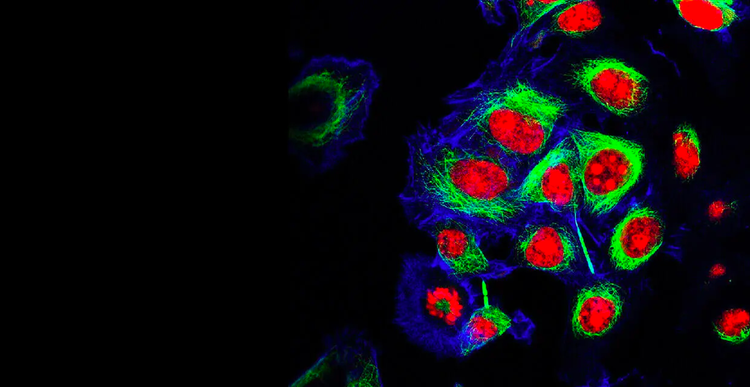
Our cell line development customers accelerate time to market for life saving monoclonal antibodies (mAbs) and genomic medicine, as well as perform groundbreaking research in cell and gene therapies, genetic engineering, personalized and precision medicine, synthetic biology, RNA and DNA-based vaccines, and so much more – made possible by our innovative technology and rich expertise.
Drug Screening
For every drug that makes it to the finish line, another nine don’t succeed. This alarming failure rate can be traced to reliance on 2D cell cultures that don’t closely mimic complex human biology, often leading to inaccurate predictions of a drug’s potential and extended drug development timelines.
A critical step in the drug discovery process, drug screening and toxicity assessment uncover the effects of potentially life-saving compounds. Shifting to cell-based testing allows multiple chemicals to be tested rapidly and better represents human biology.





