
Application Note
High throughput cardiotoxicity assays using stem cell-derived cardiomyocytes
- Earlier prediction of compound toxicity and efficacy
- Robust, high throughput, biologically relevant assays
- Fast, simplified data analysis for compound prioritization
Introduction
Cardiac toxicity is responsible for a large percentage of new drugs that fail in clinical trials. The development of highly predictive in vitro assays suitable for highthroughput screening is critical to improve the inefficiencies and high costs associated with cardiac safety compound failure. Traditional methods for characterizing cardiotoxic compounds are labor-intensive and slow. Manual patch clamping and automated electrophysiology are limited to the analysis of single channels on individual cells, and have high costs and low throughput. Other, higher throughput methods require the exporting of data into additional, often complex, software for analysis, or time-consuming manual analysis of the data. With ScreenWorks® Peak Pro™ Software running on the FLIPR® Tetra System, you can quickly and easily characterize cardiotoxic compounds using stem cell-derived cardiomyocytes. Human cardiomyocytes derived from stem cell sources can greatly accelerate the development of new chemical entities and improve drug safety by offering more biologically relevant cell-based models than those presently available. Using cardiomyocytes with ScreenWorks Peak Pro Software and the FLIPR Tetra System gives you an integrated solution that provides a robust, high-throughput method for predicting toxicity and efficacy earlier in the drug discovery process. You can fail or further monitor specific compounds sooner, and better prioritize the most promising leads to take forward into pre-clinical studies.
High-throughput FLIPR Tetra System cardiac beating assay
The FLIPR Tetra System is a flexible and reliable real-time kinetic cellular assay screening system for identifying early leads against GPCR and ion channel receptors. The system can also monitor changes in intracellular Ca2+ flux associated with cardiomyocyte contractions using the FLIPR® Calcium 5 Assay Kit. The assays use induced pluripotent stem cell (iPSC) derived cardiomyocytes (provided by Cellular Dynamics International, Inc.) loaded with FLIPR Calcium 5 Assay Kit dye, and allow you to monitor compound impact on the beating cells. The FLIPR Tetra System can automatically add reagents and compounds simultaneously while reading from 96, 384, or 1536 wells, which is advantageous for cardiac beating assays because it reduces well-to-well variability caused by reading at different time points. Temporal response curves to analyze and visualize the effect of compounds on beating cells can be acquired in approximately two minutes per plate, as shown in Figure 1 (page 2).
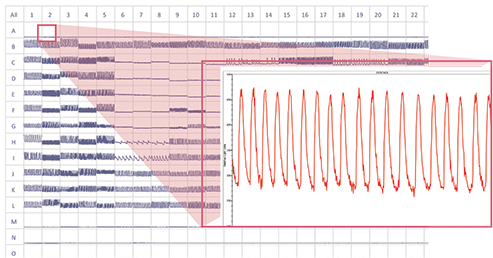
Figure 1. Beating cardiomyBeating cardiomyocytes and compound effects on response profile 384-well screenshot of concentration response curves on cardiomyocytes, increasing dose from rows K through C. Control wells in rows B and L. Measurement 5-minutes post-compound addition.
ScreenWorks Peak Pro Software automated peak analysis
Automated data analysis is a necessity to run assays on large numbers of compounds. Manual counting of 384 traces limits cell beating analysis to a few parameters, is time consuming, and subject to human error. Exporting the data to a separate program for analysis adds additional steps that compromise the quality of information or the usefulness for environments that require true automation and high throughput. ScreenWorks Peak Pro Software is integrated into the acquisition and analysis program already running on the FLIPR Tetra System. The software’s easy-to-use interface and intuitive setup provide fast and reproducible well-by-well results, with the accuracy of automated measurements.
Signal peak parameters are calculated based on a dynamic threshold and derivative analysis, and provide amplitude, time, frequency, rate, and width values, as shown in Figure 2.
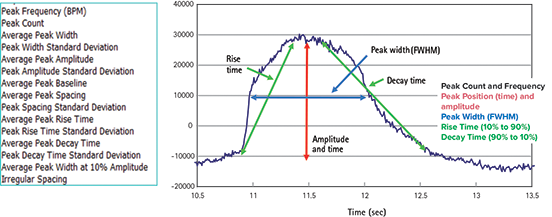
Figure 2. Software analysis output parameters Output parameters available to the user in ScreenWorks Peak Pro Software.
Figure 3 shows the benefits of ScreenWorks Peak Pro Software for identifying potentially toxic drug candidates. The cisapride data show a decrease in beat rate, increase in decay rate, and increase in peak width at 10%, all of which could have cardiac safety implications as they may be predictions of elongated QT syndrome. The ability to identify these parameters earlier in the drug discovery process could indicate the compound as a failed candidate for preclinical studies.
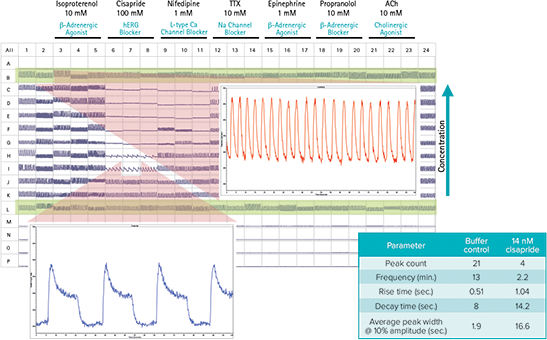
Figure 3. FLIPR Tetra System cardiomyocyte assay capabilities Output parameters available to the user in ScreenWorks Peak Pro Software.
Predictive HT cell-based assays
Development of new, more potent, and safer drugs requires in vitro systems where efficacy and safety can be tested. Positive and negative chronotropes are used in clinics to treat heart failure, tachycardia, arrhythmia and other cardiac diseases. Results from assays run on the FLIPR Tetra System show the effects of several positive and negative chronotropes, and known ion channel blockers on cardiac rates. The EC50 values shown are in the expected ranges. (See Figure 4.) This group of compounds, including a hERG channel blocker, cisapride, was analyzed using ScreenWorks Peak Pro Software. Cisapride data show a decrease in beat rate and an increase in peak width at 10% amplitude, both indicators of potentially negative cardiac effects.
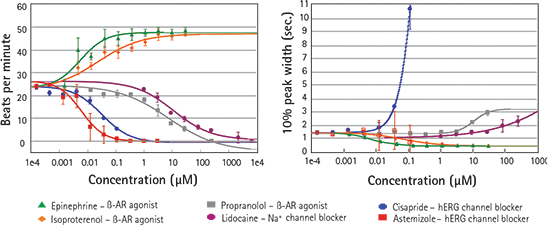
Figure 4. Effects of ß-adrenergic receptor agonists/antagonists and ion channel blockers Dose response of six different compounds as measured by the FLIPR Tetra System in Calcium 5 loaded iPSC derived cardiomyocytes.
Left: Change in frequency of contractions with dose. Right: Change in average peak width with dose.
Conclusion
The FLIPR Tetra System, together with ScreenWorks Peak Pro Software and the FLIPR Calcium 5 Assay Kit, provide a complete, integrated solution for directing your drug design by allowing you to eliminate cardiotoxic compounds and identify potential cardiac drug candidates earlier in the drug discovery process.