
AI, machine learning, and deep learning tools for cell image analysis
Artificial Intelligence in automated image analysis workflows
The recent adoption of machine learning methods, an approach to achieve artificial intelligence (AI), in image analysis is rapidly gaining momentum in many research areas. Deep learning is part of a boarder machine learning algorithm based on an artificial neural network. Deep-learning architecture successfully solved complex analysis problems in medical, pathological, and biological imaging applications.
Defining AI terms:
Artificial intelligence (AI) – A simulation of human intelligence processes by computer systems.
Machine learning (ML) – An approach to achieve AI by using algorithms to determine or predict patterns based on existing data. The machine learning algorithms then automatically infer the rules to discriminate the classes.
Deep learning (DL) - A subset of machine learning methods that use Convolutional Neural Networks (CNN) to learn input/output relationships. CNN are mathematical models represented by multiple layers of “neurons” or computational cells.
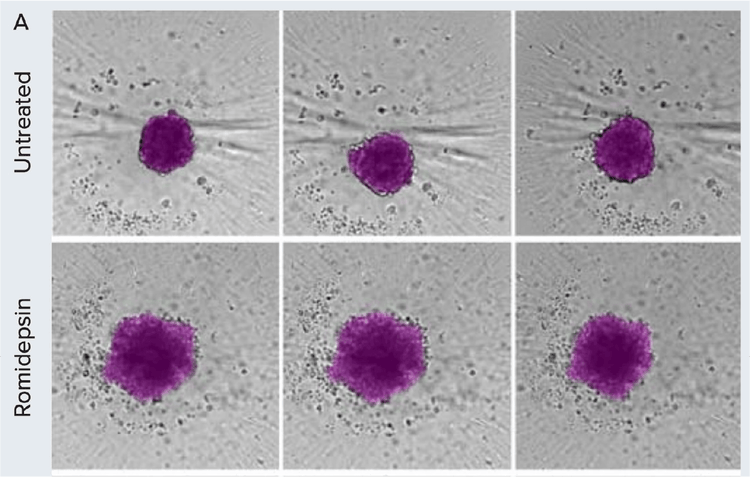
Morphometric analysis of compound treatment on patient-derived spheroids over time. A) Spheroids were monitored using brightfield imaging on days 1, 3, and 5 post-treatment. Images were segmented using SINAP in IN Carta software (magenta overlay).
Machine learning improves image segmentation and objects classification
Automated image analysis is an integral part of most high-content imaging platforms. The ability to monitor cells and organoids in real-time and then extract meaningful information depends on robust image analysis of label-free transmitted light images. Challenges associated with the analysis of brightfield images include low contrast, uneven background, and imaging artifact. A defined set of global parameters would rarely be successful in segmenting objects imaged in brightfield. Recent advances in machine learning improve image analysis workflow and enable more robust image segmentation in complex datasets.
In bioimage analysis, deep learning provides a powerful toolbox to tackle challenging image segmentation and object tracking. Conventional image analysis typically involves defining a fixed set of parameters to segment objects of interest for downstream quantification. However, these predefined parameters do not work for all experiments due to the high variability in experimental conditions. Manual adjustments to the analysis protocol are impractical because of the sheer volume of imaging data in a high-throughput environment.
To overcome these challenges, machine learning tools can be used for image segmentation and object classification to automate the image analysis workflow.
IN Carta Image Analysis Software for machine learning-based, high-throughput analysis

The IN Carta® Image Analysis Software provides an intuitive user interface that includes AI tools in the image analysis workflow. Two key components of IN Carta software that leverage machine learning are the software modules SINAP and Phenoglyphs. Deep-learning-based SINAP enables robust detection of complex objects of interest (e.g., stem cell colonies or organoids) with minimal human intervention to improve accuracy and reliability in the first step of image analysis. The analysis output includes morphological, intensity, and texture measurements.
Classification of data can also be used with machine-learning-based Phenoglyphs. Phenoglyphs takes hundreds of image descriptors extracted by SINAP and creates an optimal set of rules for grouping objects with a similar visual appearance. Both modules utilize unsupervised decisions to generate an initial result that is iteratively optimized through user input. Together, they improve the integrity and accuracy of findings through an easy-to-use, end-to-end workflow.
Apply deep learning to object segmentation with IN Carta SINAP
Automatic object segmentation of microscopy images can be challenging due to the diverse nature of the datasets. IN Carta Image Analysis Software uses SINAP, a trainable segmentation module utilizing a deep convolutional neural network learning algorithm to tackle these challenging issues.
Because SINAP uses deep learning, it can account for significant variability in sample appearance that arises from the test treatments under investigation. By ensuring that each treatment is segmented with an equivalent level of accuracy, the information extracted in this step is reliable and useful to compare treatments in subsequent analysis steps.
Overcome challenges in image segmentation with machine learning-based models:

A) Examples of different biological models are presented that are challenging for quantitative analysis. 3D spheroids grown in microcavity plates produce a shadow around each microcavity that interferes with object segmentation (arrow). 3D organoids are grown in Matrigel, which often produces non-homogenous background due to distortion from the Matrigel dome and objects beyond the imaging planes (box). iPSC grows as relatively flat cultures; as a result, the low contrast (blue arrow) and debris (yellow arrow) hampers robust image segmentation of iPSC colonies.
B) Overview of the model training workflow: Generate images for training > Train model > Test model > Repeat.
C) Main steps to create a model in IN Carta software using SINAP with example images shown. Images are annotated using labeling tools to indicate the objects of interest and background. The annotated image representing ground truth is added to the training set. In the training step, a model is created based on the most-suitable existing model and user-specified annotations. In the example shown, it is necessary to correct the segmentation mask (step 3), repeating steps 1–3.
Apply machine learning to object classification with IN Carta Phenoglyphs
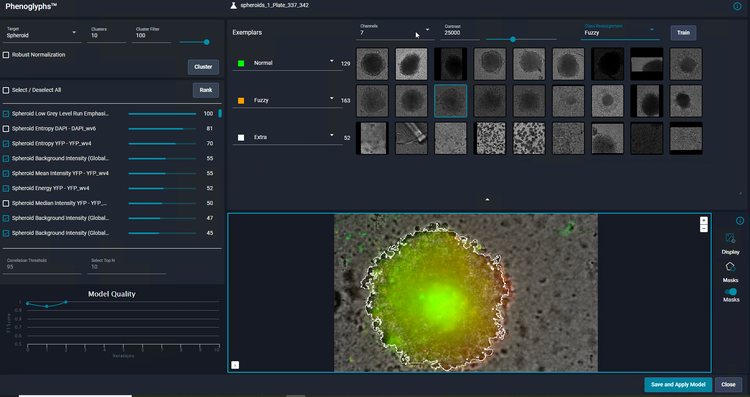
As a user, you only need to review and provide input on a small number of examples for each class before the Phenoglyphs module applies the model to the entire dataset. This approach minimizes the need for user input at the first step of a class assignment, thus saving considerable time.
Machine learning-based applications
Traditional image analysis methods can be incredibly intricate and time-consuming when performed manually or even semi-automatically. There’s always the possibility of human error and bias due to the task’s complex and highly detailed nature. When you add to this the repetitive, lengthy, and often laborious nature of the workflow, there comes the opportunity to apply machine learning.
Learn more about how IN Carta’s deep learning software paired with the ImageXpress HCS.ai system helped remove person-to-person variation, human error, and bias, thereby improving data quality and confidence and optimizing workflow and efficiency.