
Application Note
Assessing mitochondrial toxicity on the SpectraMax i3 Multi-Mode Detection Platform with SpectraMax MiniMax 300 Imaging Cytometer
- Analyze cellular images without nuclear counterstaining with StainFree Technology
- Measure mitochondrial staining on a per-cell basis with image analysis features in SoftMax Pro Software
- Obtain critical mitochondrial toxicity results faster
Introduction
A key part of drug screening programs is monitoring effects of candidate drugs on mitochondrial health. Mitochondrial activity can be readily analyzed using fluorescent dyes available in a variety of colors. These are often used in conjunction with a nuclear counterstain, such as DAPI, which enables identification of all cells in the image. Once the cells have been identified via their nuclear stain in one fluorescence channel of an imaging system, their mitochondrial activity can be measured on a per-cell basis as fluorescence in a second channel corresponding to the specific mitochondrial dye used. However, nuclear counterstaining adds an extra series of cumbersome steps to the experimental workflow and delays results. StainFree™ Technology from Molecular Devices frees researchers from the inconvenience of nuclear counterstaining by identifying cells imaged with the transmitted light channel of the SpectraMax® MiniMax™ 300 Imaging Cytometer.
This application highlight shows how the SpectraMax® i3 Multi-Mode Detection Platform can be used to assess mitochondrial toxicity with cellular imaging. Valinomycin, an ionophore antibiotic, was used to disrupt mitochondrial function in PC12 cells, and MitoTracker Deep Red FM was used to monitor the effects. Analyses using StainFree and fluorescent nuclear dye to identify the cells were compared and results evaluated using SoftMax® Pro Software. The SpectraMax MiniMax 300 Imaging Cytometer and SoftMax Pro Software provide a complete solution for image acquisition and analysis for rapid quantitation of mitochondrial activity in response to potentially toxic compounds.
Cell-based assay setup
PC12 rat pheochromocytoma cells were cultured in F-12K medium with 2.5% fetal bovine serum, 15% horse serum, and 1% penicillin/streptomycin. Cells were seeded at 10,000 cells per well at 100 µL per well, in 96-well black-wall, clear-bottom plates. Cells were allowed to attach and grow overnight.
The day after seeding, valinomycin was added to cells in the microplate in a 1:2 dilution series from 1 µM down to 1 nM. Plates were incubated for 24 hours at 37°C. Upon completion of compound incubation, culture medium plus compound in the wells was replaced with a 100 nM solution of MitoTracker Deep Red FM (MTDR) diluted in culture medium. This reagent labels active mitochondria in live cells and fluoresces at a far-red wavelength that is detected by the red fluorescent channel of the MiniMax cytometer.
Cells were incubated for 30 minutes, and then the stain was removed. Cells were then fixed with 4% paraformaldehyde. Nuclei of the fixed cells were counter-stained with EarlyTox Dead Green Dye (Molecular Devices cat. #R8216). Fixation is not required for StainFree analysis, but it was done for all cells in this experiment for comparative purposes. Cells were imaged in the transmitted light (TL) channel of the MiniMax cytometer to obtain cell counts and assess morphology, and in the red fluorescent channel to measure mitochondrial activity. Cells were also imaged in the green channel to identify the nuclei of individual cells and perform a comparative analysis alongside StainFree cell counts.
Using SoftMax Pro Software, two different image analyses were set up. In the first, individual cells were identified using StainFree analysis, which allows identification of individual cells in the transmitted light images without the need for fluorescent nuclear staining. For each cell identified, red fluorescent signal representing mitochondrial staining was measured and captured as an output parameter in the software.
A second analysis was done to confirm that results obtained with StainFree were comparable to results obtained using a nuclear stain. Here, fluorescently stained cell nuclei were identified in the green fluorescence imaging channel. Since the resulting nuclear identification masks do not include the cytoplasmic area where mitochondrial fluorescent staining occurs, the masks were extended or ‘grown’ outward by 4 pixels, or 7.5 µm. This enabled red mitochondrial fluorescence to be quantified and reported by the software for the cytoplasmic area around the nucleus.
For each analysis case, mitochondrial response to compound treatment was graphed as average mitochondrial fluorescence per cell vs. compound concentration, and IC50 values were calculated automatically by the software.
Results
PC12 cells showed a concentrationdependent reduction in mitochondrial labeling when treated with valinomycin. Whether the images were analyzed using nuclear fluorescent dye or StainFree Technology to identify individual cells, fluorescent mitochondrial staining could be quantified on a per-cell basis and used to assess mitochondrial response to compound treatment. In Figure 1, StainFree analysis is used to identify cells. Purple masks indicate individual cells.
Figure 1. StainFree analysis of PC12 cells
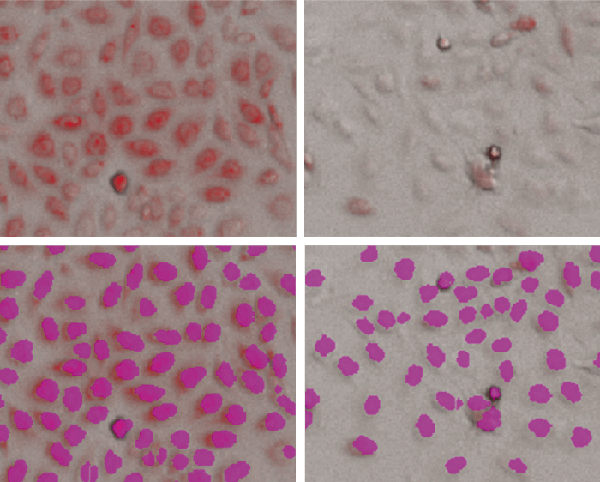
Top row, transmitted light images overlaid with red fluorescent (mitochondrial) images. Bottom row, StainFree analysis showing individual cells (purple masks) identified by the software. Left, untreated cells; right, cells treated with 1 µM valinomycin.
Figure 2 shows analysis using green fluorescently stained nuclei to identify individual cells. In this case, purple masks indicate the nuclei. The masks are grown an additional four pixels to capture more area for analysis of mitochondrial signal per cell. The lighter border around the mask indicates this extended area, whose size is user definable.
Figure 2. Fluorescent analysis of PC12 cells
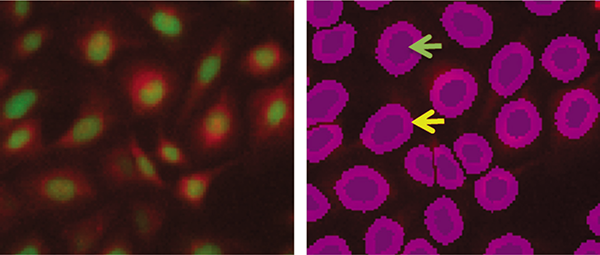
Left, green fluorescent (nuclear) images overlaid with red fluorescent (mitochondrial) images. Right, individual nuclei identified by fluorescence analysis, shown as darker purple masks (green arrow). The lighter borders indicate an extension of the object mask to capture more cellular area for analysis (yellow arrow).
Using either analysis method, the resulting IC50 values are the same (Figure 3). StainFree analysis offers a convenient alternative to time-consuming, expensive nuclear counterstaining.
Figure 3. IC50 curves for PC12 cells treated with valinomycin

Concentration-dependent response curves are shown for cells analyzed using StainFree Technology (blue) or fluorescent nuclear staining (green). Average mitochondrial fluorescence per cell is graphed vs. concentration of valinomycin. Nearly identical curves were obtained with both analysis methods, and in both cases the IC50 value was 57 nM.
Conclusion
When a fluorescent cellular marker, such as a mitochondrial dye, needs to be quantified on a per-cell basis, it is necessary to first identify individual cells. Then the fluorescent marker signal associated with each cell can be quantified. There are different ways to identify individual cells. One is to label the cells’ nuclei with a fluorescent dye, which may require fixation of the cells. A more rapid, less expensive, and less toxic way is to use StainFree Technology, which eliminates the need for fixation or dyes.
In this application highlight we have shown that StainFree Technology and the SpectraMax MiniMax 300 Imaging Cytometer can be used to first identify individual cells and then further analyze them for mitochondrial marker staining, whose intensity changes with compound treatment. The resulting concentration response curves are nearly identical to those obtained using nuclear dyes. StainFree Technology allows analysis to be performed on unfixed cells, without the need for a nuclear counterstain like DAPI. This speeds the time to results and enables a more versatile workflow in which cells can be kept alive for additional experiments.
Compatible with these Molecular Devices systems
SpectraMax i3 Platform with SpectraMax MiniMax 300 Imaging Cytometer
Learn more about SpectraMax i3x Multi-Mode Detection Platform >>