
Application Note
Alternatives to DAPI staining: imaging and counting live cells
- Enable live cell counting while maximizing cell viability with StainFree Technology
- Eliminate fixation steps required prior to staining
- Save time and money by avoiding costly staining procedures
Introduction
DAPI (4’, 6-diamidino-2-phenylindole) is a fluorescent dye often used to stain nuclear DNA. It is employed in imaging experiments such as fluorescent microscopy, chromosome spreads, FACS, and cell-based assays [1-4]. However, excitation with UV results in a photoconversion of DAPI that leads to detection of DAPI fluorescence in the FITC/GFP channel of an imaging system, causing errors in interpretation of results [5, 6]. DAPI also requires cells to be fixed for maximal staining. In this highlight, we show some alternatives to DAPI staining, including StainFree Technology, which requires no staining at all and can be used with live cells.
Molecular Devices SpectraMax® i3 Multi-Mode Microplate Platform with SpectraMax® MiniMax™ 300 Imaging Cytometer uses patented StainFree™ Cell Detection Technology to quantify cells. This label-free method uses transmitted light imaging and sophisticated software to identify individual cells or cell populations. Users can 'teach’ the software to identify cells through a supervised machine learning algorithm. This allows scientists to count cells and monitor cell growth without harming cells or losing time and money on costly staining procedures.
Another alternative to DAPI is Molecular Devices EarlyTox™ Live Red Dye. This cell-permeant red-fluorescent dye stains the nuclear DNA of all cells in culture, regardless of viability, and can be used as a substitute for DAPI in imaging assays where nuclear staining is desired. With excitation at 622 nm and peak emission at 645 nm, Live Red Dye does not interfere with the FITC channel in imaging assays.
We performed a cell-counting experiment to demonstrate performance with StainFree Technology and Live Red Dye methods vs. DAPI staining. Both alternatives to DAPI offer users the advantage of counting live cells without the need for time-consuming fixation steps. StainFree Technology frees users from the constraints of staining protocols and also enables them to use the cells for additional downstream analyses.
Methods
CHO-K1 cells were seeded into a 96-well plate at densities from 20,000 down to 300 cells per well, with serial two-fold dilutions, and allowed to attach and grow overnight at 37°C. After incubation, a subset of the wells was fixed with 4% paraformaldehyde in phosphate-buffered saline (PBS) for 30 minutes at 37°C. Following fixation, cells were washed with 1X PBS two times and then incubated in 0.1% Triton X-100 for 5 minutes. Cells were washed twice with PBS and stained with 2.9 µM DAPI in PBS for 30 minutes. After staining, cells were washed three times with PBS.
The remaining cells were left alive and stained with Live Red Dye. A stock solution of the dye was diluted 1:2000 in PBS and added to cells. After a 30-minute incubation, live cells were imaged with the transmitted light and red fluorescent (Cy5) channels on the MiniMax cytometer. StainFree analysis and nuclear counts were performed with SoftMax Pro Software. Figure 1 shows the time saved using StainFree or Live Red Dye.
For comparison, live (red-stained) and fixed (DAPI-stained) cells were imaged with the Cy5 and DAPI channels in the ImageXpress® Micro XLS Widefield High-Content Analysis System. In order to compare cell counts on both imaging systems, a region of interest (ROI) was applied to the images acquired with the MiniMax cytometer to match the area imaged on the ImageXpress Micro XLS system. Graphs were created using SoftMax Pro Software.
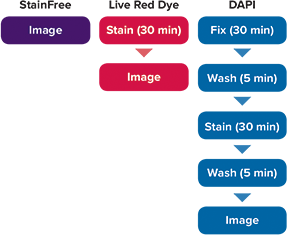
Figure 1. Workflow for cell analysis with StainFree Technology vs. Live Red Dye vs. DAPI. The StainFree workflow saves about 70 minutes compared to fixation and staining with DAPI. Moreover, cells analyzed using StainFree Technology remain fully viable and can be used in additional assays.
Results
Cells stained with Live Red Dye and imaged on red fluorescent and transmitted light channels of the MiniMax cytometer are shown in Figure 2. For the red fluorescent images, a predefined ‘Nuclei’ setting in SoftMax Pro Software was used to accurately identify stained nuclei. For cells imaged in the transmitted light channel, StainFree Technology was used to identify cells. The predefined setting ‘CellsA’ in the software achieved the most precise results for these CHO cells. StainFree cell counts aligned very closely with fluorescent nuclear counts (Figure 2, graph), demonstrating that the MiniMax cytometer and SoftMax Pro Software can eliminate the need for a nuclear stain to count cells accurately.
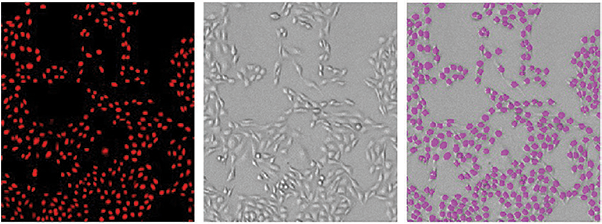
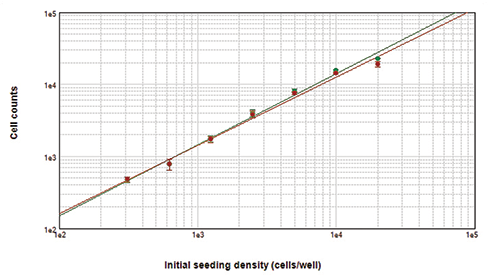
Figure 2. Cells counted with the MiniMax cytometer. Cells stained with Live Red Dye were imaged in the red fluorescent (top left) or transmitted light (top middle) channel. StainFree cell counts (top right, purple masks) correlate very closely to cell counts based on red nuclear staining, as shown in the graph (green circles, StainFree counts; red circles, red nuclear counts). The r2 values for both curves were 0.99.
DAPI-stained cells were imaged and counted using the ImageXpress Micro XLS system with MetaXpress® Software. For comparison, Live Red Dye-stained cells were imaged on the same system using the Cy5 channel. Images of stained cells are shown in Figure 3. Cell counts with both dyes agreed very closely (Figure 3, graph). From this comparison, we confirm that cell counts using DAPI are equivalent to those using StainFree Technology.
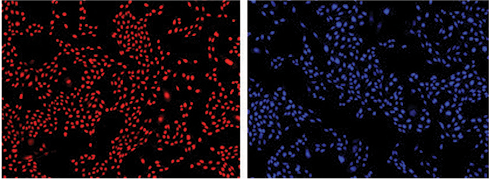
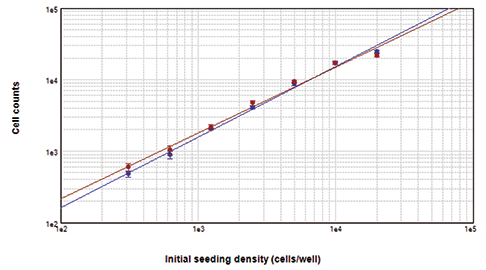
Figure 3. Cells counted on the ImageXpress XLS system. Cells stained with Live Red Dye (upper left) or DAPI (upper right) were imaged and close correlation of counts obtained with both dyes was demonstrated by graphing in SoftMax Pro Software (bottom). For both curves, r2 was 0.99.
Conclusion
With equivalent cell counts, StainFree Technology and Live Red Dye are just as effective for cell counting as DAPI, which requires fixation prior to staining. StainFree cell counting is the best choice for maximizing cell viability and minimizing time spent on cell handling as it requires neither fluorescent dyes nor fixation. For cases where a nuclear dye is desired, e.g. to monitor nuclear size or staining levels, Live Red Dye can be used in place of DAPI but does not require fixation. It also will not interfere with green fluorescent imaging as DAPI has been shown to do. Using either StainFree Technology or Live Red Dye allows greater versatility for live cellbased assays, and greater confidence in experimental results.
References
- https://www.lifetechnologies.com/order/catalog/product/D1306
- Otto F. 1990. DAPI staining of fixed cells for high-resolution flow cytometry of nuclear DNA. Methods in Cell Biology 33: 105-110.
- Kapuscinski J. 1995. DAPI: a DNA-specific fluorescent probe. Biotechnic & Histochemistry 70(5): 220-233.
- Chazotte B. 2011. Labeling nuclear DNA using DAPI. Cold Spring Harb Protoc; 2011; doi:10.1101/pdb.prot5556.
- North AJ. 2006. Seeing is believing? A beginners’ guide to practical pitfalls in image acquisition. J. Cell Biology 172(1): 9-18.
- Jez M, Bas T, Veber M, Kosir A, Dominko T, Page R, Rozman P. 2012. The hazards of DAPI photoconversion: effects of dye, mounting media and fixative, and how to minimize the problem. Histochem. Cell Biol. 131(1): 195-204.
Learn more about SpectraMax MiniMax 300 Imaging Cytometer >>