
Application Note
Image slide-based samples on the SpectraMax MiniMax cytometer
- Acquire high-quality images from slide-based samples
- Identify and analyze a variety of different tissue types
- Create custom plate dimensions for microscope slides, petri dishes and other non-microplate formats
- Accessible to non-imaging experts with intuitive SoftMax Pro Software
Introduction
Biological samples ranging from bacterial smears to tissue cross sections are commonly studied on microscope slides. Using bright-field and fluorescence imaging, researchers can study cytological or morphological changes in their slide-based samples. Slide microscopy has traditionally required microscope systems, but microplate reader systems have begun to evolve the capability to perform slide-based imaging.
The SpectraMax® i3x Multi-Mode Microplate Reader with the SpectraMax® MiniMax™ 300 Imaging Cytometer can be used to image slides using a specialized slide adapter. Equipped with transmitted light (TL), green (Ex/Em: 460/541) and red (625/713) fluorescence channels, the MiniMax cytometer with SoftMax® Pro Software can be used to image and analyze a variety of tissue-based samples.
This application note demonstrates how the MiniMax cytometer can acquire high-quality images and data from slide-based samples.
Materials
- SpectraMax i3x Multi-Mode Microplate Reader (Molecular Devices cat. #i3x)
- SpectraMax MiniMax 300 Imaging Cytometer (Molecular Devices cat. #5024062)
- Slide adapter (Molecular Devices cat. #1-5200-1337)
- FluoCells Prepared Slide #1 (ThermoFisher cat. #F36924)
Methods
A slide adapter was used to hold the microscope slide samples (Figure 1). A “custom plate setting” was created in SoftMax Pro Software to accommodate the dimensions of the slide adapter by editing a current plate type and saving it under a different name. To create a custom plate, click on “Settings”, select the “Plate Type” tab, and click “Edit Plate” (the 96-well plate format was used in this example). Input the dimensions as shown in Figure 2 to accommodate the slide-based adapter and save it under a custom name. This custom plate setting can then be used for future slide-based imaging experiments.
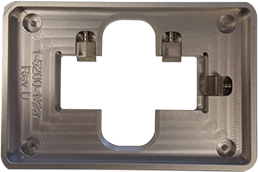

Figure 1. Slide adapter. The slide adapter has been specifically designed with an SBS footprint to enable the imaging of microscope slides in a microplate reader.
The FluoCells® prepared slide was placed onto the slide adapter and imaged using the same custom plate setting created previously. The acquisition settings used are shown in Table 1. The FluoCells slide contained fluorescently-stained bovine pulmonary arterial endothelial (BPAE) cells. SoftMax Pro Software was used to calculate “% cell coverage”. The “Field Analysis” setting using the 541 wavelength was selected, and the pre-made “Cells” setting was used to identify cell-confluent regions. These identified regions are shown with a purple overlay as seen in Figure 3. The calculated % of cell coverage is displayed in the associated plate section and the data tables in SoftMax Pro Software.
Transmitted light (TL)
541 nm (green fluorescence)
713 nm (red fluorescence)
TL exposure: 7 ms
TL focus adjustment:
-460 μm*
541 exposure: 200 ms
541 focus adjustment:
-410 μm*
713 exposure: 1600 ms
713 focus adjustment:
-440 μm*
Analysis type: Field analysis
Wavelength for finding objects: 541
*Focus adjustment values are indicated for the slide type used here. Other slides may require different focus adjustment values.
Table 1. Acquisition and analysis settings for the FluoCells prepared slides.
Tibia cross-section slides were imaged using the transmitted light (TL) setting (Figure 4). Samples were imaged in a 384-well format, and only the proximal portions of the tibia samples are shown. SoftMax Pro’s StainFree Analysis was used to calculate the area (μm2) of identified bone tissue. Underneath the Image Analysis tab, the “Field Analysis” setting using the TL wavelength was selected. The pre-made “Cells” setting was used to identify regions occupied by tibia sample as shown with a purple overlay. The area is calculated and displayed in corresponding plate sections and the data tables in SoftMax Pro Software. In this example, we display the total bone area underneath the images in Figure 4.
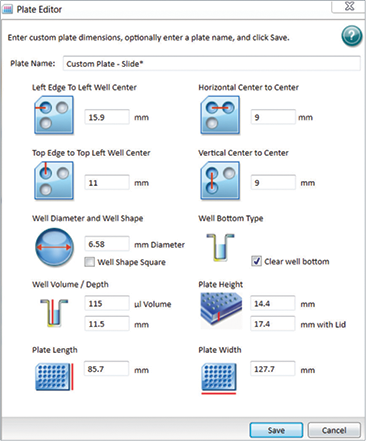
Figure 2. Custom plate dimensions for the slide adapter. A custom plate setting was created in SoftMax Pro Software to accommodate the slide adapter using the dimensions shown above.
Results
The prepared slide containing fluorescently stained BPAE cells was imaged using the slide adapter and settings shown in Figure 1 and 2. We were able to acquire high-quality fluorescent images from these slides and SoftMax Pro was able to calculate cell confluency based on the green fluorescence (Figure 3).
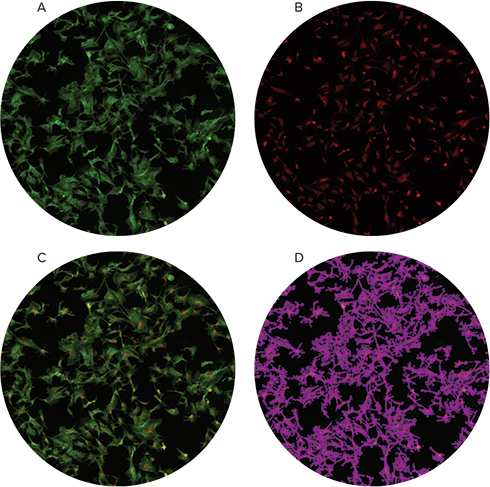
Figure 3. FluoCells prepared slide imaged and analyzed. A pre-made slide containing BPAE1 cells stained with Alexa Fluor® 488 Phalloidin and MitoTracker® Red CMXRos was imaged using the MiniMax cytometer. Green fluorescence channel images (A) and red fluorescence channel images (B) were overlaid (C), and cell confluency as determined by Field Analysis of the green fluorescence channel was identified in panel D.
Additionally, the MiniMax cytometer was able to image tissue sections using the same slide adapter and the custom plate definition created in SoftMax Pro Software. In the example below, tibia bone samples were imaged and the approximate sample area is calculated using the StainFree analysis (Figure 4). The total bone area can be calculated by adding together the calculated area of each well containing bone sample, which can then be used to approximate bone growth.
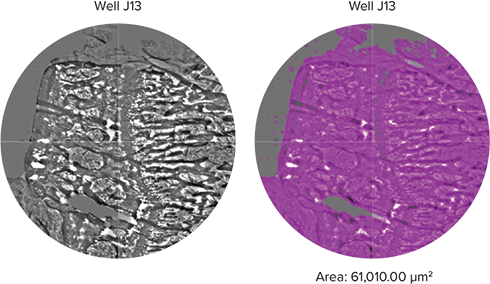
Figure 4. Tibia cross section. A cross section of human tibia bone was placed onto a slide and imaged using the TL setting. A portion of the tibia cross section is shown on the left, and the object mask is shown on the right.
Conclusion
With the addition of the plate editor and the slide adapter, the SpectraMax MiniMax 300 Imaging Cytometer can be used to acquire high-quality images from disparate sample types such as microscope slides. The SpectraMax i3x microplate reader along with the MiniMax cytometer can analyze these samples in numerous ways increasing the scientist’s research capabilities.
Learn more about SpectraMax MiniMax 300 Imaging Cytometer >>