
Application Note
Easy picking with the QPix 400: multiple selection modalities, wide range of microorganisms
- Fast—up to 30,000 colonies/day, >98% efficiency
- Efficient transfer—agar level sensor for automated picking pin height adjustment
- Optimized—organism-specific pins
- Targeted—quantitative, userdefined selection criteria
- Smart—intelligent colony selection software
Introduction
Whether you’re trying to discover the next generation of antibiotics, selecting clones for sequencing, or turning micro algae into biofuel production factories, chances are you’ll need to screen thousands, maybe even millions of colonies to find the best ones. Again and again. And not just screen, but plate and replicate.
Automated colony pickers can simplify and speed these laborious processes, but what if you need to screen instead of select, choosing colonies based on morphology instead of just the ability to grow on restrictive media? What if you’re working with a microbe other than E. coli? With the QPix™ 400 series of microbial colony pickers, you can do all that—morphologybased colony picking, plating, and plate replication of bacteria, fungi, algae, phage, and yeast cells.
The QPix 400 colony pickers support a wide variety of microorganisms and multiple selection modalities, including fluorescence intensity, blue/white selection, size and proximity, zone of inhibition—you set the selection parameters and the instrument does the work.
Colony selection workflow
Whatever your selection modality, setting selection parameters for any QPix 400 colony picker follows the same general workflow—open the QPix software and define your colony selection parameters, such as size, compactness, and other morphological features. For Zone of Inhibition detection, set your parameters in the Zone of Inhibition Detection Module. Colonies are detected by white light and, if desired, further selected using fluorescence intensity.
Colony selection in white light
Selection of colonies visualized in white light is a typical first step for most studies using the QPix colony picker. In this example, Streptomyces venezuelae was plated, incubated at 37°C overnight, and colonies selected in white light. The intelligent software module analyzes the images, and colonies that meet the userdefined selection criteria are outlined in yellow and then picked (Figure 1). Colonies that do not meet the selection criteria are highlighted in red.
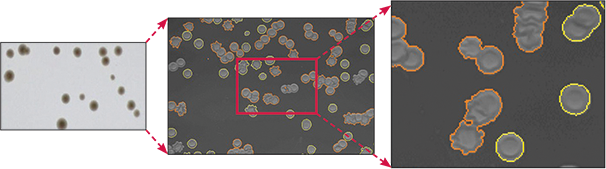
Figure 1: S. venezuelae plated (left panel). The intelligent software module detects desired colonies based on user-defined criteria (middle and right panels, yellow-outlined colonies), and rejects colonies that do not meet the selection metrics (middle and right panels, red-outlined colonies).
Colony selection by fluorescence
With fluorescence-based selection downstream processing time can be significantly reduced through earlier selection of high-value targets (Figure 2). Through the use of appropriate fluorescent markers, morphological and functional screens can be combined in the automated selection phase, saving time and resources by requiring fewer colonies to be further screened and characterized in a downstream functional assay.
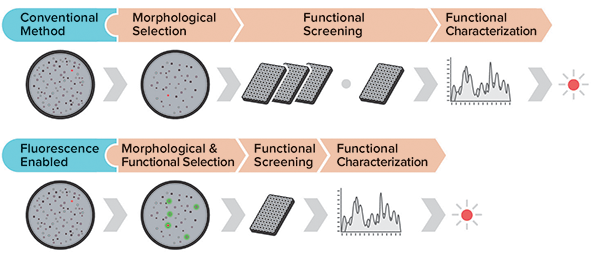
Figure 2: Fluorescent selection saves screening time by focusing downstream assays and screening on high-value targets.
An example of fluorescence-based functional selection can be seen in this experiment where Nile Red, a lipophilic fluorescent dye, was used to select high lipid-producing strains of Rhodococcus opacus PD630 for biofuel production. Colonies with higher lipid content show correspondingly higher fluorescence intensities. For this experiment parameters such as size, diameter, and compactness were tuned for optimal colony selection, and a mean fluorescence intensity threshold of greater than 50,000 was set to select (i.e. gate) colonies demonstrating high fluorescence as a result of high lipid accumulation (Figure 3).
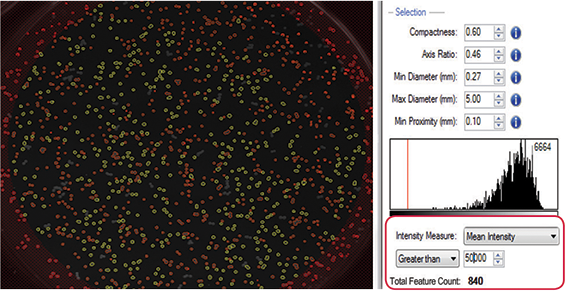
Figure 3: Fluorescence-based quantification that is reflective of lipid accumulation enables objective colony selection on QPix 400 Series.
Lipid accumulation levels were further confirmed post picking. Following overnight growth in liquid media, cultures were stained with a lipophilic bright green fluorescent dye, BODIPY 505/515 (Life Technologies), at a concentration of 0.5 μg/mL, and fluorescence measurements were recorded on SpectraMax® M5 Multi- Mode Microplate Reader. A high degree of fluorescence was confirmed in high lipid-accumulating colonies that were originally selected and picked using the QPix 420 System, while background levels of fluorescence were exhibited by the negative control E. coli colonies (Figure 4, red curve).
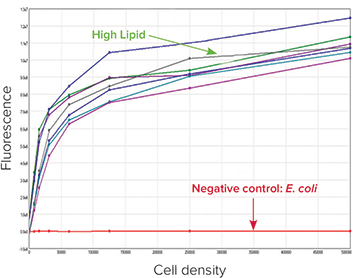
Figure 4: Fluorescence reading of high lipid producing colonies and negative control group stained with BODIPY 505/515 lipophilic fluorescent dye on the SpectraMax M5 Multi-Mode Microplate Reader. High lipid producing colonies exhibited high fluorescence (RFU) values and in contrast the negative colonies exhibited a flat-line signal as depicted in red.
Colony selection by blue/white color screening
Selecting colonies by blue or white color enables the use of the widely-used LacZ reporter system for selecting recombinant/ non-recombinant clones. For this example white light images taken on the QPix 420 System were analyzed using the easyto-use QPix™ Software 2.0. Blue (Figure 5A) or white (Figure 5B) colonies were automatically identified and selected separately using the built-in Auto Select feature (Figure 5C). By adjusting the histogram threshold and defining colony selection criteria such as compactness, axis ratio, diameter, and proximity, you can further optimize your selection.
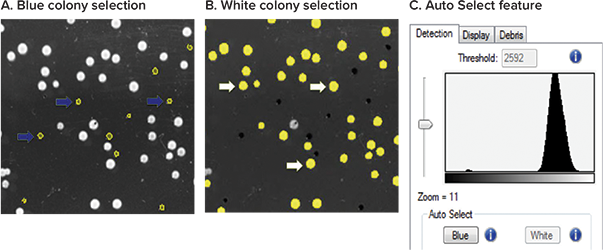
Figure 5: (A) Example of blue colonies selected with Auto Select Blue are indicated with blue arrows. (B) Example of white colonies selected with Auto Select White are indicated with white arrows. (C) Flexibility of the software allows manual adjustment of the intensity threshold to optimize results, such as to select for powered blue colonies.
Selection of colonies secreting zones of inhibition
For a library-based approach to screening and selection of antibiotic producing organisms, the agar plate-based zone of inhibition or clearing zone detection assay is the method of choice. Colonies providing antimicrobial activity are distinguished from the rest of the population by secreting clearing zones where bacterial growth is inhibited due to secretion of antimicrobial compounds. The diameter of the clearing zone is typically proportional to the amount of antimicrobial compound produced. Thus, high value strains are those that generate the largest clearing zones.
With the Zone of Inihibition Detection module in QPix Software 2.0 or higher, the size of each antimicrobial-producing colony and the size of the clearing zone produced by that colony can both be reliable quantified. You can select colonies that produce the largest clearing zones (Figure 6).
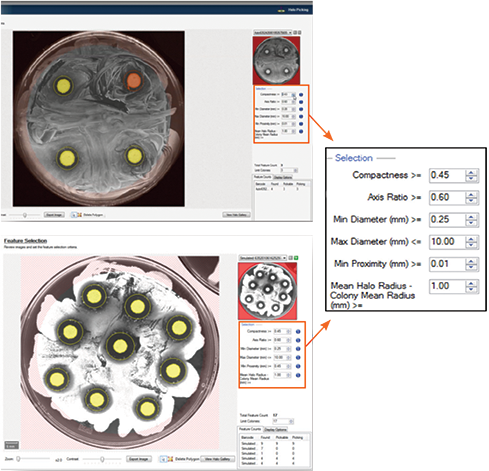
Figure 6: The Zone of Inhibition Detection module in QPix Software 2.0 or higher can identify the size of each colony and corresponding clearing zones. Colonies producing clearing zones are selected based on user-defined selection criteria.
To demonstrate the utility of automated zone of inhibition detection for librarybased screening or adaptive evolution studies, QTrays spotted with a library of microbial cultures were screened and selected. From the 48 colonies grown on QTrays, three colonies producing zones of inhibition were reliably detected by the QPix Software Zone of Inhibition Detection module (Figure 7), selected, and picked.
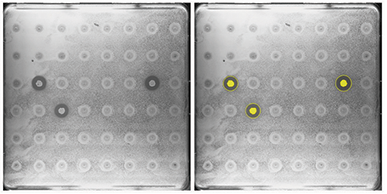

Figure 7: Library based high-throughput colony screening of zone of inhibition producers for antibiotics discovery. The Zone of Inhibition Detection module in QPix Software 2.0 or higher enables reliable detection and selection of colonies producing clearing zones. Selected colonies producing corresponding clearing zones are highlighted in yellow.
This approach is amenable to highthroughput implementation aimed to screen, identify, and pick colonies producing clearing zones from a large microbial library.
Picking more than E. coli
Repetitive failure of colony transfer can result in project delays, wasted biomaterials, or the loss of valuable clones. To ensure optimal colony transfer for a diverse range of microorganisms, Molecular Devices offers a unique portfolio of picking pins that come in a variety of shapes, sizes, and textures. Each organism-specific pin is designed to meet the designated microbial colony’s shape, stickiness, viscosity, or other characteristic that can impact picking efficiency.
In addition, a proprietary agar height sensor automatically determines the optimal picking height on-the-fly, thus enabling optimal transfer and outgrowth of biological materials.
Together, proper pin selection and automated agar height sensing enhance microbial colony transfer efficiency by as much as 40% as shown in an example dataset (Figure 8).
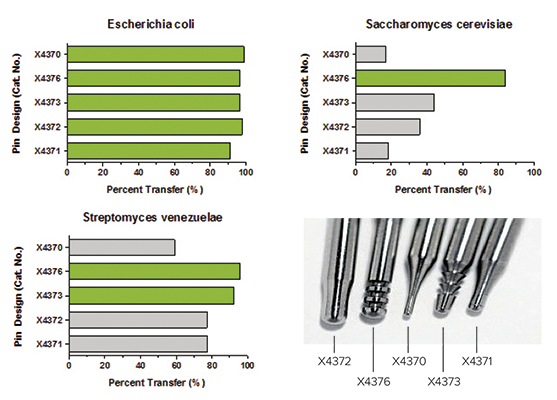
Figure 8: Colony transfer efficiencies can vary widely according to the pairings between microorganisms and colony-picking pins. A representative dataset is shown for E. coli, Saccharomyces cerevisiae, and S. venezuelae.
The QPix software is also designed to support selection of a wide range of microbial species, as a colony detection algorithm optimized for E. coli may not work optimally when applied to yeast, algae, or other microorganisms with different phenotypic attributes. The QPix Software offers experimental flexibility by automatically identifying and selecting colonies produced from a wide range of microorganisms (Figure 9). Furthermore, the ability to custom define parameters such as shape, size, and proximity to neighboring objects ensures efficient and tailored selection of any colony forming microorganism, and microorganism-specific pins ensure efficient colony transfer.
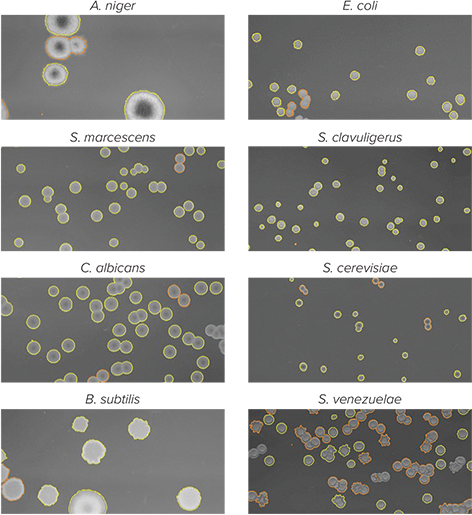
Figure 9: Morphologically distinct colonies are identified by QPix colony detection algorithms based on user-defined parameters: compactness, axis ratio, size, and proximity. Colonies surrounded by yellow border are selected for picking based on the user criteria, while colonies bordered with red fail to meet user-defined criteria and are excluded from picking.
Summary
Through sophisticated algorithms, easy-touse software with customizable selection criteria, and organism-specific algorithms and accessories, the QPix 400 series of colony pickers is a unique solution where automation is in synergy with life science needs. Colony picking precision and speed, hardware solutions optimized for different organisms, and robust software algorithms offer the flexibility and power to handle any number of screening projects. With the QPix 400, selecting the right colony is easy pickings.
The Molecular Devices QPix team continues to add new application notes to our library www.moleculardevices.com/qpix-resources