
Cell Counter
At the Cell Counter: MCF-7 Cells
Since October is breast cancer awareness month, we decided to highlight the MCF-7 breast cancer cell line. Derived in 1970 from a nun named Sister Catherine Frances (Helen Marion) Mallon, this cell line was named after the Michigan Cancer Foundation, where it was generated. MCF-7 is one of the few breast cancer cell lines to express substantial levels of estrogen receptor (ER) alpha, thus it offers researchers a valuable model system for the study of ER-positive breast cancers. Type ‘MCF-7 cells’ into PubMed and you will find over 20,000 references to this remarkable cell line.
View StainFree Cell Detection Webinar
Download StainFree Cell Detection Application Note
Download eBook: Count Cells Like a Pro
Figure 1: Images obtained using SpectraMax MiniMax cytometer
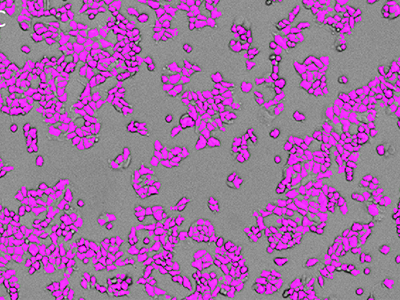
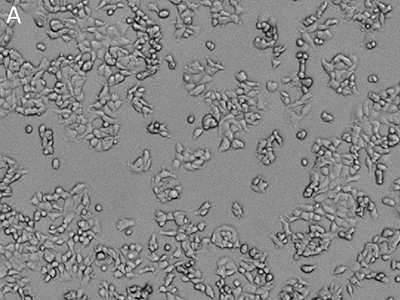
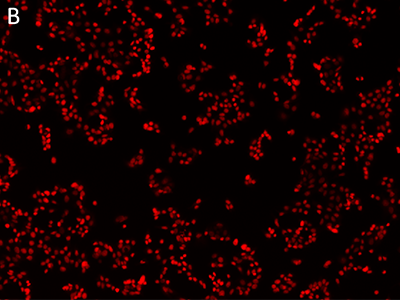

MCF-7 cells were stained with EarlyTox™ Live Red Dye and then imaged using the SpectraMax MiniMax cytometer. Cells were identified with a user-defined custom setting. A, transmitted light image; B, nuclei stained with EarlyTox Live Red Dye; C, cells identified using StainFree analysis (purple masks show cells identified by the software).
Figure 2: StainFree cell counts vs. fluorescent cell counts
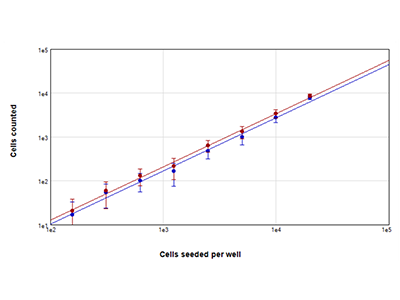
MCF-7 cells were seeded at densities ranging from 156 to 20,000 cells per well, and nuclei were stained using EarlyTox Live Red dye. Images were acquired from 4 sites per well, and a region of interest in the middle of the wells was selected for analysis. Cells were then counted by using either the StainFree method (blue plot) or by counting red-fluorescent nuclei (red plot).
Figure 3: EarlyTox Live/Dead Assay
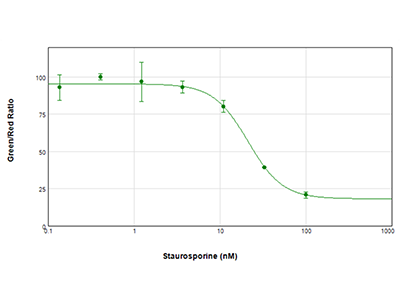
MCF-7 cells were treated with serial dilutions of staurosporine for 24 hours and then assayed using Molecular Devices EarlyTox Live/Dead Assay Kit. Live cells were stained with Calcein AM (green dye), and dead cells were stained with an ethidium homodimer (red dye). The plate was read on the SpectraMax i3x Multi-Mode Microplate Reader. Results were plotted as ratio of green/red vs. compound concentration, and an IC50 value of 21.2 nM was calculated using SoftMax Pro Software.
Tip:
MCF-7 cells have a polygonal morphology with irregular dimensions, and they have a habit of clumping into large aggregates. For StainFree cell counting, use ‘Create New Setting’ and draw slightly within the cell borders using the yellow drawing tool. Use the blue drawing tool to mark areas outside cell boundaries. Alternatively, you can use the ‘CellsC’ predefined setting, but it may not be quite as accurate.
MCF-7 Cells Analysis Toolkit
- SpectraMax ® i3x Multi-Mode Microplate Detection Platform
- SpectraMax ® MiniMax™ 300 Imaging Cytometer
- SoftMax ® Pro Software
Transmitted light (TL)
713 nm (red fluorescence)
TL exposure: 7 ms
TL focus adjustment: -10 µm
713 exposure: 6 ms
713 focus adjustment: 0 µm
Analysis type: Discrete Object Analysis
Wavelength for finding objects: TL (StainFree) or 713 (red nuclei)
About StainFree Cell Detection Technology
Imaging cell-based assays typically requires the use of fluorescent probes that can be toxic to living cells or may only function in fixed cells. A label-free method for analyzing cell counts and cell confluence enables researchers to quantitatively monitor cell proliferation and health without time-consuming workflows that may disrupt cell viability.
The SpectraMax i3 Multi-Mode Microplate Platform with MiniMax 300 Imaging Cytometer uses unique, patent-pending StainFree Cell Detection Technology that allows you to perform cell proliferation, cytotoxicity, and other assays without nuclear stains like DAPI, which intercalates with DNA, or live cell dyes that are actually toxic to cells in the long term.