
Application Note
Nucleic acid quantitation and analysis using the QuickDrop Spectrophotometer
- Minimal sample usage as little as 0.5 μL
- Accurate DNA Quantitation from 1.0 ng/μL to 2,500 ng/μL
- LCD touchscreen for stand-alone experimentation and data analysis
- Kinetic and wave scanning capabilities for cuvette reads
Introduction
Spectrophotometry is a well-established technique used to quantitate and analyze biological substances. Of these substances, nucleic acids are one of the most routinely measured in life science laboratories. Determining the concentration and purity of these samples is extremely important for many downstream experiments such as PCR, qPCR, sequencing, and DNA microarrays.
Nucleic acids absorb ultraviolet light mainly at 260 nm, and their concentrations can be calculated by applying the Beer-Lambert law with their associated extinction coefficient and sample path length. First, ultraviolet light at 260 nm is directed into the sample, and as this light passes through the sample, a photodetector on the other side measures how much light was absorbed. By comparing the results to a reference (usually the sample diluent), one can quantitate a sample’s nucleic acid concentration.
Sample purity is an important parameter in nucleotide quantitation. Although it is not the most accurate method for determining sample purity, the A260/ A280 and A260/A230 ratios can be used as a rough estimate of protein and chemical contamination.
The SpectraMax® QuickDrop™ Spectrophotometer is a versatile ultraviolet-visible (UV-Vis) micro-volume spectrophotometer exceptional at analyzing nucleic acid samples. It contains a 0.5 mm micro-volume port, cuvette port, and built-in LCD touchscreen allowing the user to perform various experiments all on one system.
In this application note, we demonstrate how the QuickDrop spectrophotometer can both quantitatively (concentration) and qualitatively (sample purity) analyze nucleic acid samples with great accuracy and consistency.
Materials and methods
- QuickDrop UV-Vis Spectrophotometer (Molecular Devices cat. #QUICKDROP))
- UltraPure ™ Calf Thymus DNA Solution (Thermo Fisher cat. #15633019)
- RNA Control 250 (Thermo Fisher cat. #AM7155)
- 10 mm Far UV Quartz Cuvettes (Starna Cells cat. #9-Q-10)
Sample carryover
Sample carryover was assessed by alternating measurements on the microvolume port with calf-thymus DNA and ultra-pure water. Ultra-pure water was used as a reference. The micro-volume port was wiped with a lint-free wipe after each read.
Standard curve linearity
A two-fold dilution series starting from 2500 ng/μL of double-stranded DNA (dsDNA) was prepared using ultra-pure water. Ultra-pure water was used as a blank, and each sample concentration was read three times on the 0.5-mm micro-volume port. Using the preprogrammed DNA quantification method, absorbance was measured over wavelengths ranging from 230 nm to 320 nm. The DNA quantification method automatically calculated the dsDNA concentration based on the equation shown below where A260 and A320 refer to the absorbance measured at 260 nm and 320 nm respectively.
Concentration in μg/mL = ( A260 –A320 ) x dilution factor x 50 μg/mL
The QuickDrop automatically performs these calculations and reports the concentration to users. Afterwards, data were graphed using SoftMax® Pro Software and the SoftMax Pro Import Feature. A log-log curve fit was applied to demonstrate the linearity of the standard curve.
Sample volume comparison
Calf thymus DNA was prepared by diluting stock calf thymus DNA with ultra-pure water. 0.5 μL, 1.0 μL and 2.0 μL of samples were tested on the micro-volume port (n = 5). Ultra-pure water was used as a reference. Reference volumes were similar to the sample volumes tested.
RNA validation
ThermoFisher’s RNA Control 250 was used to compare the QuickDrop’s spectrophotometer performance to that of the NanoDrop™ Spectrophotometer. The storage solution provided with the kit was used as the reference. Both the reference sample and the RNA Control were assayed on the micro-volume port. Results were compared to the specifications provided in the RNA control kit.
Contaminant detection
Sample DNA was contaminated with a phenol solution resulting in a final concentration of 50 μM phenol in the solution. Contaminated sample and control were then analyzed in the cuvette port using the “Wavescan” feature. This feature incrementally checks the sample’s absorbance over a range of wavelengths. Results were graphed using Microsoft® Excel.
Results
Sample carryover
Sample carryover experiments were tested on the QuickDrop’s micro-volume port. Results in Figure 1 and Table 1 demonstrated that with a simple wipe of a lint-free wipe, there was no significant sample carryover in the subsequent reads.
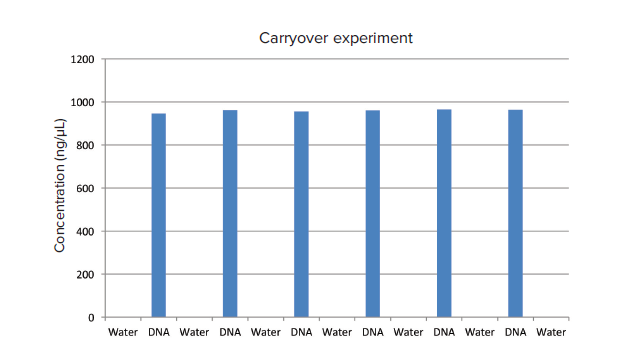
Figure 1. Sample carryover. Alternating measurements of calf thymus DNA and water were performed on the micro-volume port. Data were graphed on Excel. The data comprising this graph are shown in Table 1.
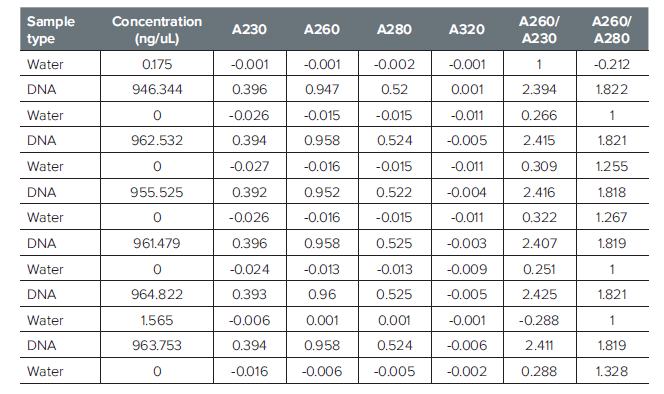
Table 1: Data comprising the “Sample carryover” experiment in Figure 1.
Standard curve linearity
In Figure 2, the QuickDrop demonstrated a linear relationship between the DNA concentration and the dilution factor. From this experiment, we determined that the 1 ng/μL DNA was the lowest measureable concentration for this QuickDrop.
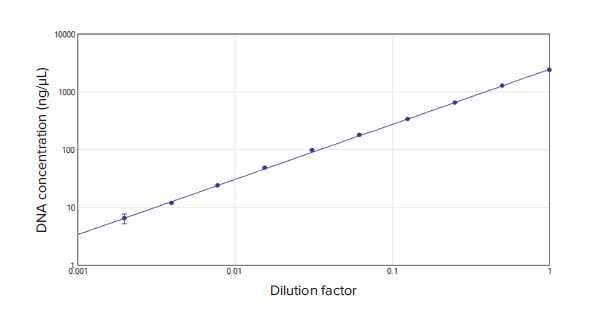
Figure 2. Calculated DNA concentration vs. dilution factor. A 2-fold dilution series of calf thymus DNA starting from 2500 ng/μL shows the relationship between the dilution factor and the DNA concentration reported by the QuickDrop. The curve demonstrated excellent linearity with a r2 value of 1.000.
Sample volume comparison
Three different volumes of sample DNA were assayed on the QuickDrop’s micro-volume port. The calculated sample DNA concentration was consistent despite the volume difference (Figure 3). For best results, we recommend using a 2-μL volume as it was the easiest to pipette.
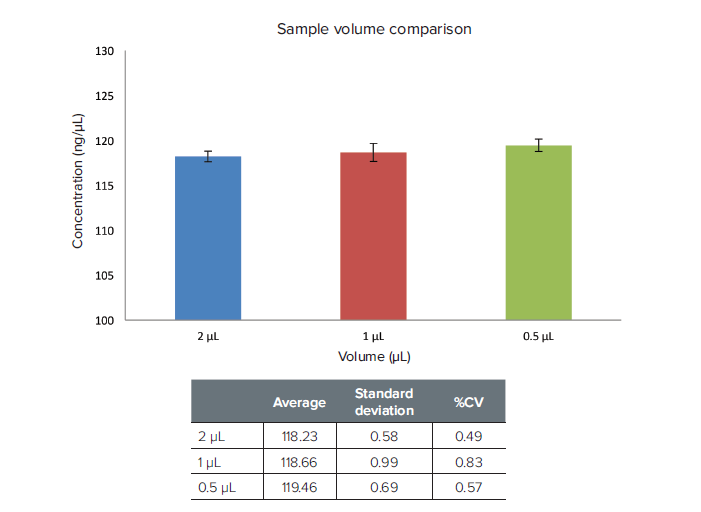
Figure 3. Volume reproducibility. 2 μL, 1 μL, and 0.5 μL volumes of calf-thymus DNA were read on the micro-volume port. Different volumes demonstrated very similar calculated concentrations (n = 5). Standard Deviation and %CV were very similar for all sample volumes.
RNA validation
An RNA 250 control was measured to demonstrate the QuickDrop’s ability to measure RNA. The measured concentration was within the accepted limits provided by the control kit (250 ± 5 ng/μL).
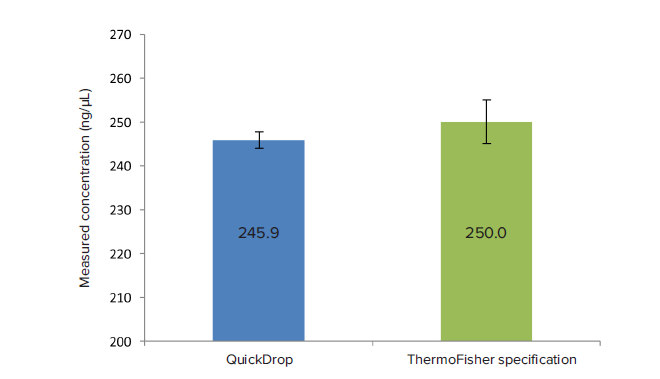
Figure 4. RNA validation experiment. 250 ng/μL RNA Control was measured on the QuickDrop to compare results to ThermoFisher’s NanoDrop validation results (n = 4). The measured RNA concentration (245.9 ± 1.9 ng/μL) falls within ThermoFisher’s specified range (250.0 ± 5.0 ng/μL).
Contaminant detection
Using the Wavescan function, contaminants can be identified in samples due to the increase in absorbance at 230 nm compared to a control (Figure 5). This function can be applied to other types of contaminants found in the DNA purification process such as guanidine thiocyanate. The effective wavelength range of the QuickDrop extends from 190 nm to 1100 nm allowing for a comprehensive chemical analysis.
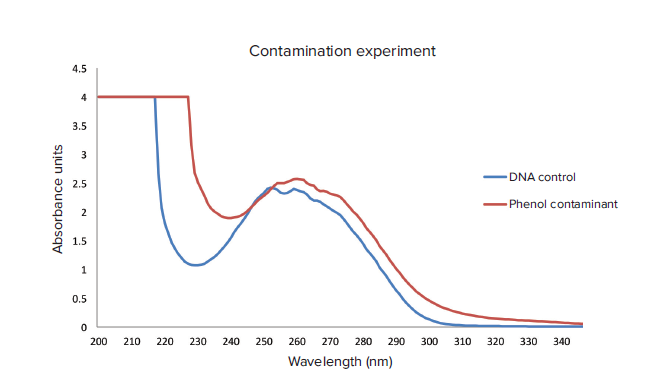
Figure 5. Contaminant experiment. Phenol-contaminated samples and an uncontaminated control were assayed using the Wavescan function. Wavescan traces were graphed and overlaid using Microsoft Excel. There is a definitive increase in absorbance at the 230 nm mark.
Conclusion
The QuickDrop’s built-in cuvette port and micro-volume sample port allow for both sensitive quantitation and analysis of nucleic acid samples. As shown above, as little as 0.5 μL of sample can be read on the micro-volume port with great consistency. Also, it has a wide detection range (1 ng/μL to 2500 ng/μL) and low sample carryover.
Finally, its built-in LCD touchscreen provides researchers with important information such as sample concentration and purity (Figure 6). All these features, along with its small footprint, make the QuickDrop Spectrophotometer an excellent choice for nucleic acid quantitation and analysis in any lab environment.

Figure 6. Example of QuickDrop’s data display. Relevant parameters are calculated and reported on the QuickDrop’s touchscreen.
Learn more about SpectraMax QuickDrop Micro-Volume Spectrophotometer >>
简介
分光光度测定法是一项定量和分析生物成分的成熟技术。其中,核酸是生物实验室最常检测的生物成分之一。确定这些样品的浓度和纯度对许多下游实验至关重要,例如PCR、qPCR测序和DNA基因芯片。
核酸主要吸收260nm下的紫外光,其浓度可以应用比尔-朗伯特定律通过它们的相关消光系数和样品光程计算出来。首先,260nm的紫外光直接照射样品,并且穿过样品,而另一边的光电检测器则测定有多少光被吸收。通过对照参比(一般是样品稀释液),可以定量样品中的核酸浓度。
样品纯度是核苷酸定量的一个重要指标。尽管不是确定纯度最准确的方法,A260/A280和A260/A230依然可以用来粗略估计蛋白和化学成分的污染程度。
SpectraMax® QuickDrop™分光光度计是 一款多用途的紫外-可见(UV-Vis)超微量分 光光度计,尤其是在分析核酸样品方面。 它包含一个0.5 mm超微量上样孔、比色皿 插槽和内置的LCD触摸屏,使用者可以在 这一个系统上进行多种实验。
在这一应用指南中,我们展示了QuickDrop分光光度计是如何以高准确度和高一致性来定量(浓度)和定性(样品纯度)分析核酸样品的。
材料和方法
- QuickDrop 紫外-可见分光光度计 (Molecular Devices cat. #QUICKDROP)
- UltraPure™ 小牛胸腺DNA溶液 (Thermo Fisher cat. #15633019)
- RNA Control 250 (Thermo Fisher cat. #AM7155)
- 10 mm 远紫外石英比色皿 (Starna Cells cat. #9-Q-10)
样品交叉污染
样品交叉污染通过使用超微量上样孔交替检测小牛胸腺DNA和超纯水进行评价。超纯水作为参比。超微量孔在每次读数完成后用不起毛的纸擦拭干净。
标准曲线的线性
溶于超纯水中起始浓度为2500 ng/µL的双 链DNA(dsDNA)经两倍系列稀释。超纯水 作为空白,每个样品浓度都在0.5-mm超 微量上样孔中读3次。利用预设好的DNA 定量方法,从230nm到320nm波长下检测 器吸光度值。DNA定量法自动计算出 dsDNA浓度,基于计算方程及以下260nm 和320nm分别检测出的吸光度值。
浓度µg/mL = ( A260 –A320 ) x 稀释因子 x 50 µg/mL
QuickDrop自动运行这些计算并报告浓度 给使用者。之后,数据可以使用SoftMax® Pro软件和SoftMax Pro导入功能绘制图 表。双对数曲线拟合用于显示标准曲线的 线性。
样品体积比较
小牛胸腺DNA采用超纯水稀释,分别是0.5 µL, 1.0 µL和2.0 µL,在超微量上样孔中检 测(n = 5)。超纯水作为参比。参比体积与样 品体积相同。
RNA 验证
ThermoFisher的RNA质控250用于比较QuickDrop分光光度计和NanoDrop™分光光度计的性能表现。试剂盒中提供的储存液作为参比。参比样品和RNA质控品均于超微量上样孔中检测。所得结果与RNA质控试剂盒说明书中的参数做比较。
污染物检测
样品DNA被50 µM苯酚溶液污染。污染的 样品和质控品在比色皿模块中使用“波长 扫描”功能检测。这一功能在整个波长范 围内递增的检测样品吸光度。结果使用 Microsoft® Excel绘制曲线。
结果
样品交叉污染
样品交叉污染在QuickDrop超微量上样孔中检测。图1和表1中的结果显示经过不起毛的简单擦拭后,后续实验中没有明显的样品交叉污染。

图 1. 样品交叉污染。 小牛胸腺DNA和水在超微量孔中的交替测定。数据在Excel中绘制图表。图表中的数据如表1。

表 1:图 1“样品交叉污染” 实验中所包含数据。
标准曲线线性
在图2中,QuickDrop展示出DNA浓度和 稀释因子之间的线性关系。从这个实验 中,我们可以得出QuickDrop的最低检测 浓度为1 ng/µL DNA。

图 2. 计算的DNA浓度 vs 稀释因子。 。起始浓度为2500 ng/µL小牛胸腺DNA经2倍系列稀释所显 示出的稀释因子和DNA浓度之间的关系,结果来源于QuickDrop。这一曲线展示出r2 值为1.000 的线性关系。
样品体积比较
3个不同体积DNA样品在QuickDrop超微量上样孔中进行检测。计算出的样品DNA浓度在不同体积下都是一致的(图3)。为了得到更好的结果,我们建议使用2-µL体积,因为其更易加样。

图 3. 体积再现性。 2 µL、1 µL和0.5 µL体积的小牛胸腺DNA在超微量孔中读数。不同体积显示 出非常一致的计算浓度(n = 5)。 标准差和%CV在所有样品体积下也都非常相似。
RNA验证
对RNA 250质控品的检测说明了QuickDrop完全能够用于检测RNA。所测浓度在 质控品试剂盒提供的可接受范围之内(250 ±5 ng/µL)。

图 4. RNA验证实验。 250 ng/µL RNA质控品在QuickDrop上检测,与ThermoFisher的 NanoDrop验证结果比较(n=4)。检测出的RNA浓度(245.9 ± 1.9 ng/µL)落在ThermoFisher的 指定范围之内(250.0 ± 5.0 ng/µL)。
污染物检测
使用波长扫描功能,污染物可以从样品中识别出来,通过在230nm处与对照比较,有上升的吸光度值(图5)。这一功能可以应用于DNA纯化过程中其他类型污染物的发现,如硫氰酸胍。QuickDrop的有效波长范围可从190nm到1100nm,对全面分析各类化学品都很有帮助。

图 5. 污染物实验。 苯酚污染的样品和未污染的对照利用波长扫描功能进行测定。波长扫描曲线 通过Microsoft Excel绘制并叠加。在230nm处可以看到有显著的光吸收。
总结
QuickDrop内置的比色皿模块和超微量 上样孔都能够进行非常灵敏的核酸样品 定量及分析。如以上所示,低至0.5 µL的样 品也能在超微量孔中得到一致性很高的 读数 。同时,还具有非常宽的检测范围 (1 ng/µL to 2500 ng/µL)和非常低的样品交 叉污染。
最后,内置的LCD触摸屏为研究者提供了重要信息,如样品浓度和纯度(图6)。所有这些特点,加上它小巧灵活的体积,QuickDrop分光光度计将成为在任何实验室环境下进行核酸定量和分析的理想选择。

图 6. QuickDrop数据显示举例。 相关参数可通过操作QuickDrop触摸屏计算并报告出来。