
Application Note
Quantification of spheroid growth and morphology
- Quickly evaluate 3D spheroids using simple acquisition and analysis techniques
- Evaluate population-level information to better understand compound effects
- Minimize biological interactions using label-free techniques
Andy Bashford, PhD | Field Applications Scientist | Molecular Devices
Matteo Cattaneo, PhD | Field Applications Scientist | Molecular Devices
Introduction
There is a growing trend to develop screening assays using three-dimensional (3D) cell culture. A common approach is to grow cells within an extracellular matrix allowing greater interaction with neighboring cells and more complex structure formation. Here, we describe a simple assay to assess cytotoxicity of chemotherapeutic agents on a human colorectal carcinoma cell line. This method delivers an easy and fast approach for quantifying spheroid growth and gross morphological characteristics. Structures can be detected using a fluorescent nuclear stain and analyzed using the CellReporterXpress® Image Acquisition and Analysis Software to assess Summary-level (per well) and population-level information. Label-free analysis can also be used to minimize assay preparation time or monitor growth in live cell environments.
This assay demonstrates a common approach used in preliminary live cell studies to assess 3D spheroids in an automated way using the ImageXpress® Pico Automated Cell Imaging System. This type of assay can be used to inform more complex mechanistic studies which can be completed using confocal imaging systems such as the ImageXpress® Micro Confocal High-Content Imaging System(1-3).
Materials
- HCT 116 cells (ATCC®, #CCL-247™)
- McCoy’s 5a growth media + 2 mM Glutamine + 10% Fetal Bovine Serum
- Corning® Matrigel Membrane Matrix, Phenol Red Free (Corning, #356237)
- Corning® 96-well Half Area High Content Imaging Film Bottom Microplate (Corning, #4680)
- Etoposide in DMSO; final well concentration of 3 μM and 100 μM (Merck, #E1383)
- Methanol free, ultra-pure Formaldehyde; 16% stock, diluted to 4% final well concentration (Polysciences, #18814-20)
- Hoechst 33342; 75 μg/mL stock in PBS diluted to 15 mg/mL final well concentration (Invitrogen, #H2570)
- ImageXpress Pico Automated Cell Imaging System with CellReporterXpress Image Acquisition and Analysis Software (Molecular Devices)
- MetaXpress® High-Content Image Acquisition & Analysis Software (Molecular Devices)
Methods
- HCT 116 cells were seeded in Matrigel at 4,000 cells per well in 50 μL volume of 96-well half area microplate. After allowing the gel to form, 50 μL of media was added on top of the gel and cells allowed to grow for 24 hours.
- Cells were treated with etoposide to a final concentration of 3 μM and 100 μM and allowed to incubate for 72 hours.
- Cultures were fixed with formaldehyde for eight hours at room temperature.
- Cultures were washed twice with PBS stained with Hoechst for six hours at room temperature.
- Cultures were washed with PBS and plates sealed ready for imaging.
- 3D cell cultures were imaged using the ImageXpress Pico system using the DAPI and transmitted light channels. Two sites were acquired per well using the 10X objective. Hardware-based autofocus was used to acquire two z-planes with 10 μm separation. DAPI images were analyzed using best-focus 2D projection using CellReporterXpress software.
- Label-free image analysis was completed using MetaXpress software. Images were imported into the MetaXpress software and analysis configured using the Custom Module Editor.
Results
Assess compound effects on spheroid size and number
3D spheroids could be imaged throughout the matrix using the z-stack functionality. The 10X magnification allowed a large number of spheroids to be captured per field of view. Images were compressed into 2D maximum and best-plane projection images reducing storage space and analysis time.
3D spheroids were analyzed using the DAPI nuclear stain to assess average spheroid size and number (Figure 1A). Etoposide caused a dose-dependent effect reducing the average spheroid size. Despite their smaller size, a high number of spheroids remained visible in treated wells (Figure 1B). The highest concentration of etoposide created a high level of debris and single cells, to improve accuracy these were excluded from the analyses and only the spheroids were quantified.
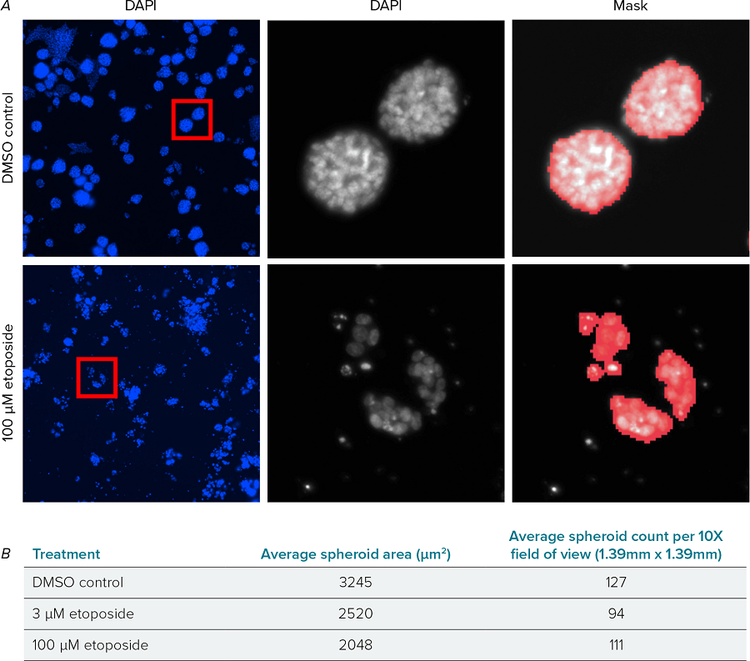
Figure 1. Analysis of DAPI labeled spheroids. A) DAPI images of DMSO control (top) and etoposide treated (bottom) wells acquired with the ImageXpress Pico system. Whole field of view 10X (left), cropped spheroids (middle) with analysis mask (right) created using CellReporterXpress software. B) Summary measurements showing average spheroid area and average number quantified per field of view for DMSO control and etoposide treated wells. Data averaged from four wells per condition, two fields of view per well.
Evaluate spheroid population using object-by-object data
Spheroid-by-spheroid data was reviewed using the CellReporterXpress software to better understand the population within each well. Population-level measurement data was also exported as .csv and further analyzed in Microsoft Excel. Visualizing the data as a box plot showed that each condition had several large spheroids which were outliers to the population (Figure 2A). Visualizing the data as a frequency plot showed that both low and high concentration etoposide caused a reduction in the number of large spheroids sized 2500–5500 μm2 (Figure 2B). Interestingly, high concentration of etoposide also caused an increase in the number of small structures sized 500–2000 μm2. This suggests that growth inhibition may be responsible for the reduction in large spheroid size but cell death and loss of spheroid integrity may play a greater role at higher compound concentrations.
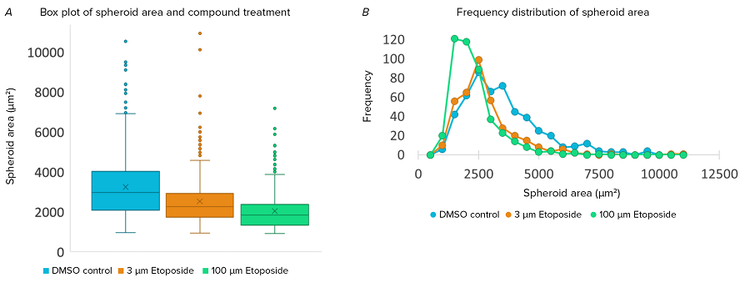
Figure 2. Population analysis of spheroid area. A) Box plot and whisker showing distribution of spheroid area. 0 μM etoposide (blue), 3 μM etoposide (orange), 100 μM etoposide (green). Data pooled from four wells per condition, two fields of view per well. B) Frequency distribution of spheroid area (500 μm bin size). Combined population from treated wells.
Label-free analysis to measure spheroid morphology
Label-free analysis of transmitted light images is an excellent way to monitor the growth of spheroids without interfering with biological factors. Transmitted light images were imported into MetaXpress software and used to mask spheroid structures. The flexibility of the Custom Module Editor allowed the exclusion of structures touching the border, in order to improve accuracy, and allowed morphological data to be extracted (Figure 3).
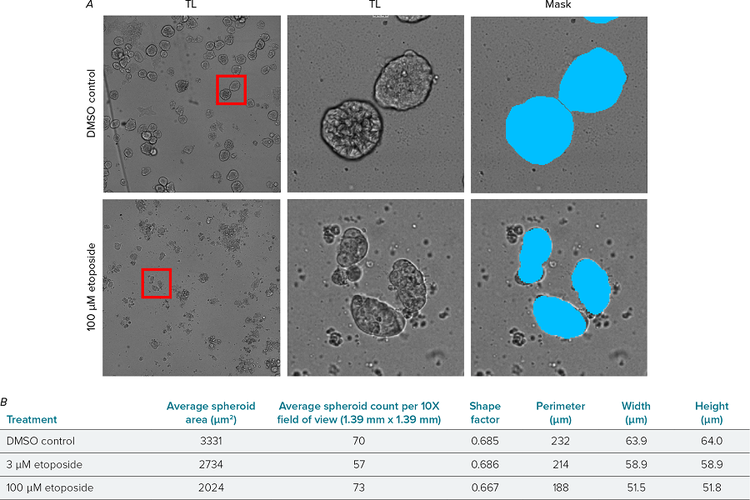
Figure 3. Label-free analysis of spheroids. A) Transmitted light (TL) images of DMSO control (top) and 100 μM etoposide treated (bottom) wells. Whole field of view 10X (left), cropped spheroids (middle) with analysis mask (right) created using the Custom Module Editor. B) Average area, spheroid count, shape factor, width, height measurements using label-free analysis. Data averaged from four wells per condition, two fields of view per well.
Average spheroid area measurements were very similar to those calculated using fluorescent marker analysis. Low concentration etoposide (3 µM) caused a reduction in average spheroid area but the shape factor remained the same as DMSO control. High concentration etoposide (100 µM) spheroids were smaller and also showed a small reduction in shape factor demonstrating that spheroids were more irregular in shape. This type of label-free analysis could remove the need for fixation and staining, making assays quicker and easier to perform in the laboratory. It could also be used in live cell cultures to track changes in growth and morphology over time.
Conclusion
Here, we demonstrate a simple method for assessing spheroid growth and morphology using fluorescence-based or label-free analyses. The assay showed a dose-dependent reduction in spheroid size in response to compound treatment. The imaging protocol is easy to set up and automate across multi-well plates and is applicable in many situations where scientists require a quick and straightforward readout from 3D cell cultures. It also shows the capabilities of label-free analyses which can be used to monitor 3D cultures in live cell experiments. This type of study is made easy with the ImageXpress Pico system using acquisition features such as z-stack, Digital Confocal* 2D on-the-fly deconvolution and ‘discontinuous time-lapse’, the latter enabling users to remove their plate from the system and return it only when needed for longer duration of live cell experiments.
The data presented suggests a shift from growth inhibition at low compound concentrations to cell death and loss of spheroid integrity at high compound concentrations. To better understand the mechanism of action, advanced experiments to analyze the volumetric 3D structure and quantify cellular components could be completed using a system such as the ImageXpress Micro Confocal system(1-3). The pairing of the two systems would allow users to match the complexity of their assays to the required assay readouts.
References
- Application Note: Multi-parameter imaging assay for measuring toxicity in a tumor model
- Application Note: High-throughput confocal imaging of spheroids for screening cancer therapeutics
- Application Note: High-content assay for morphological characterization of neuronal development in a 3D matrix using human iPSC-derived neuronal cultures
Supplementary acquisition and analysis methods
Quantification of spheroid growth and morphology—a fast and simple approach for compound screening
Plate setup
Prior to the start of the experiment, an empty microwell of the same batch to be used to grow cells was inserted into the ImageXpress Pico system and the Measure Plate Dimensions option used to configure the Well Depth, Bottom Thickness, and Plate Clearance. The remaining plate dimensions were completed according to the manufacturer’s documentation (Figure S1).
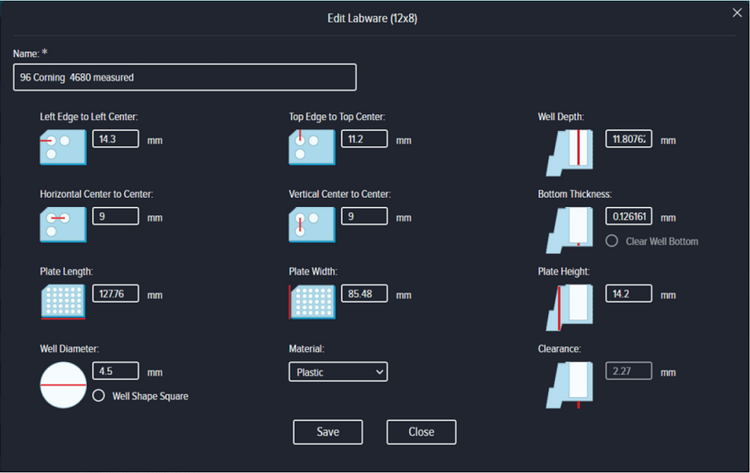
Figure S1. Plate dimensions of Corning 96 A/2 film bottom, high content microplate measuring using the CellReporterXpress software.
Acquisition
Focus method was set to ‘PlateAndWellBottom’ with a Focus offset set to 0 μm, and Exposure set to 10 ms for both DAPI and transmitted light (Figure S2). Z-stack acquisition was enabled and a 2-plane stack with Focus Step of 10 μm acquired at +10 and +20 above the bottom of the well (Figure S3). Maximum and Best Plane projection were selected for Fluorescent and transmitted light channels respectively.
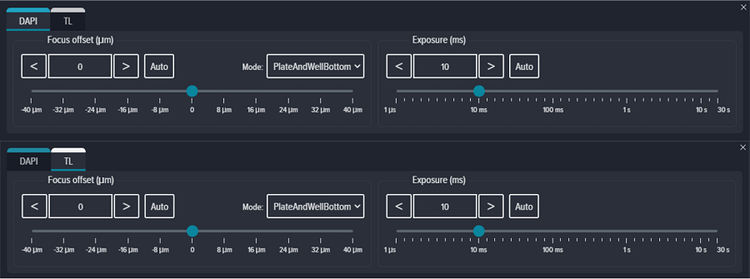
Figure S2. Plate focus offset and exposure for DAPI and TL channel.
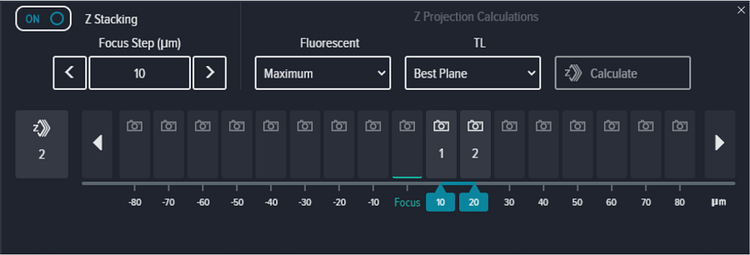
Figure S3. z-stack settings showing Maximum and Best Plane 2D projection image.
CellReporterXpress analysis
The plate was first annotated with compound and concentration using the Plate Annotation tool.
Fluorescent DAPI images were analyzed using the Cell Count analysis in CellReporterXpress software (Intensity = 100, Min Width = 30, Max Width = 100). All Summary and Cell Measurements were selected to ‘ON’ (Figure S4).
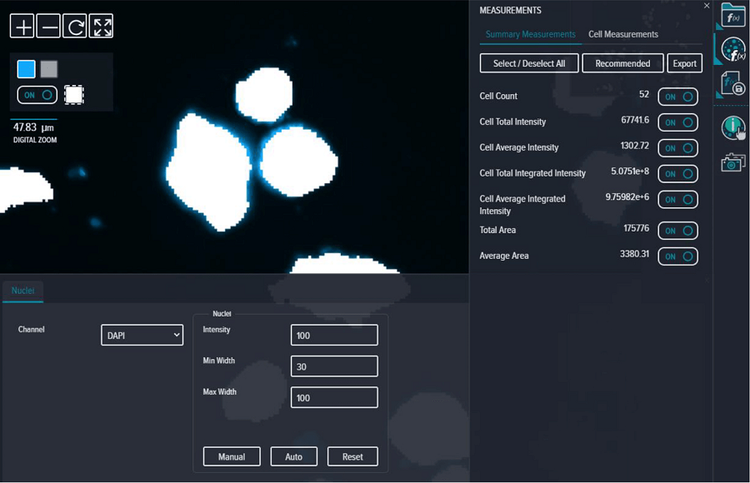
Figure S4. Analysis settings for spheroid identification using DAPI stain.
The analysis was run on the whole plate and object data was exported as a ‘.csv’ format. This data was opened in Microsoft Excel where Box Plot and Frequency distribution were used to display the population information.
MetaXpress analysis
The Export Experiment Images tool was used to export images from the CellReporterXpress software as full size ‘.tif’ format in an organized folder structure.
Images were imported into MetaXpress software using the MDCStore Xchange tool and a template recognizing the folder structure and naming convention (Figure S5, template can be provided on request).
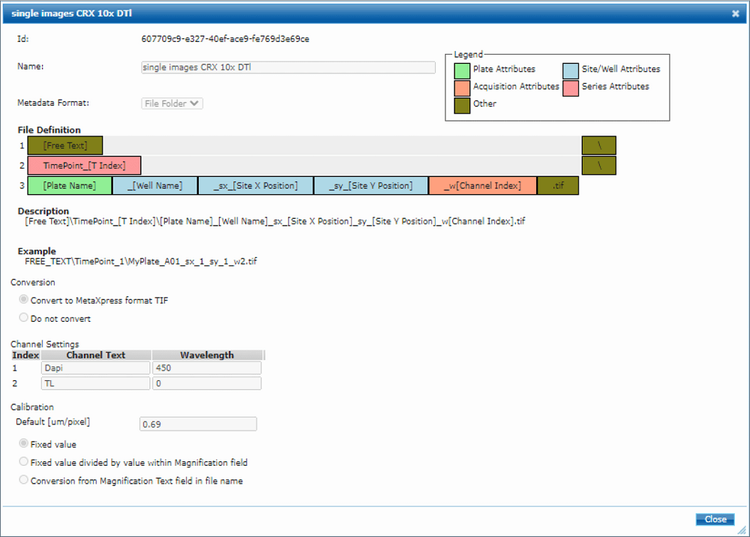
Figure S5. MDCStore Xchange template for import of images into MetaXpress software.
Transmitted light images were analyzed using the Custom Module Editor. A module was designed to recognize the structures and eliminate those touching the edge (please see below for custom module download link). Analysis was run on the whole plate.
Custom module steps
1. HBasinThreshold = 1800
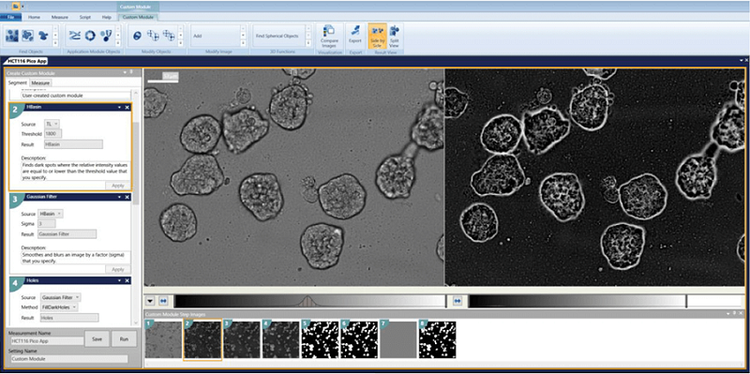
2. Gaussian Filter Sigma = 3
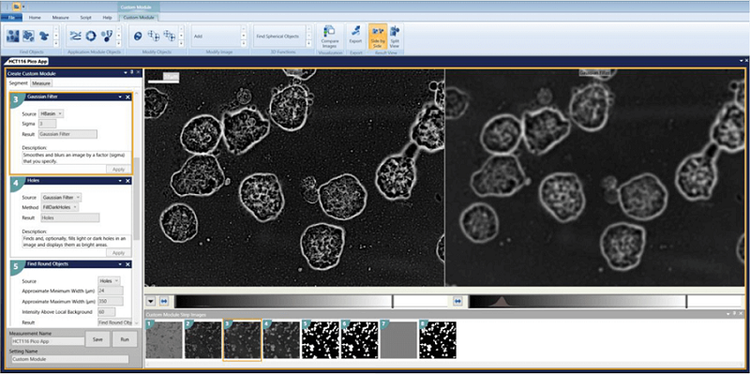
3. Holes = FillDarkHoles
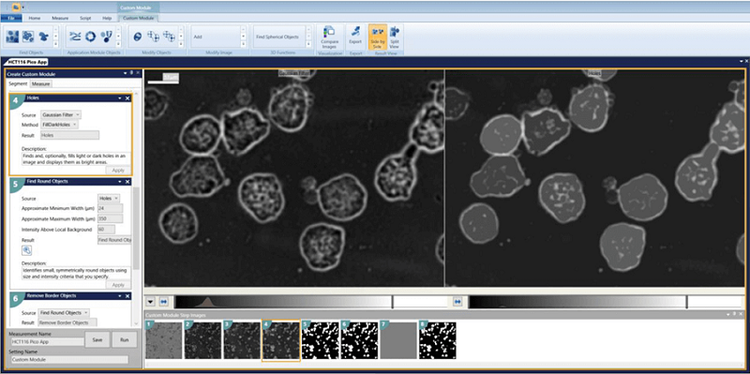
To download the custom module, please visit https://moleculardevices.box.com/v/pico-custom-module
4. Find Round Objects Minimum width = 24 Maximum width = 350 Intensity above local background = 60
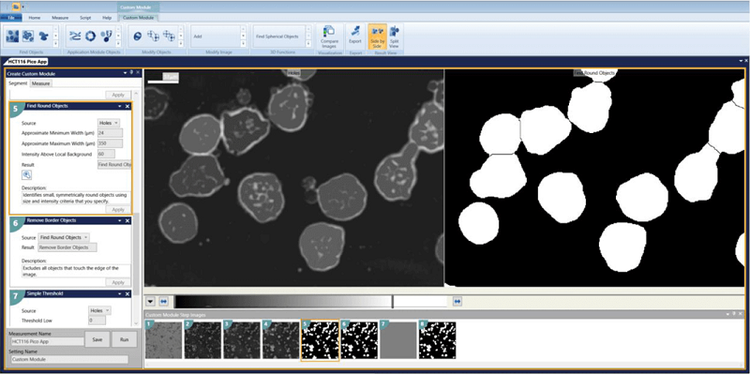
5. Remove Border Objects
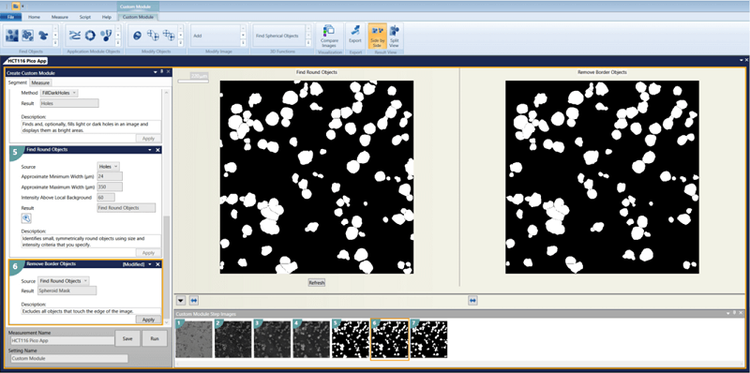
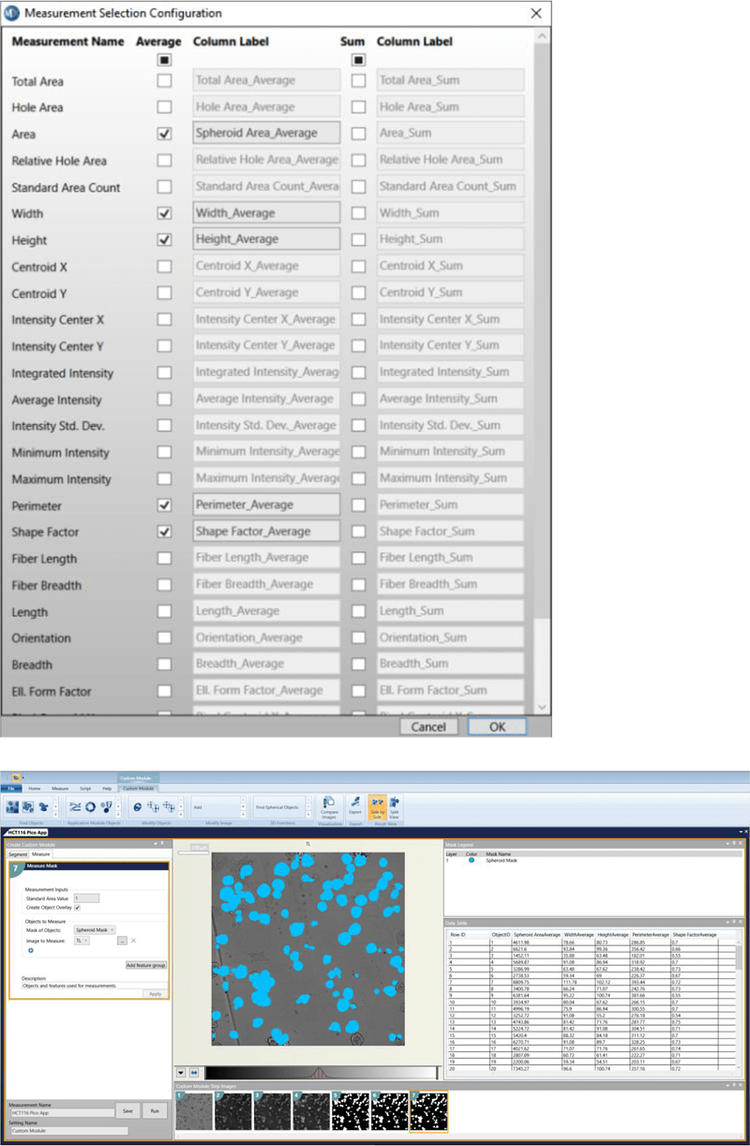
6. Measurements to select: Area, Width, Height, Perimeter, Shape Factor
*ImageXpress Pico Digital Confocal uses AutoQuant 2D Real Time Deconvolution